Volume 16, Number 4—April 2010
Research
Phylogenetic Analysis of Enterohemorrhagic Escherichia coli O157, Germany, 1987–2008
Figure
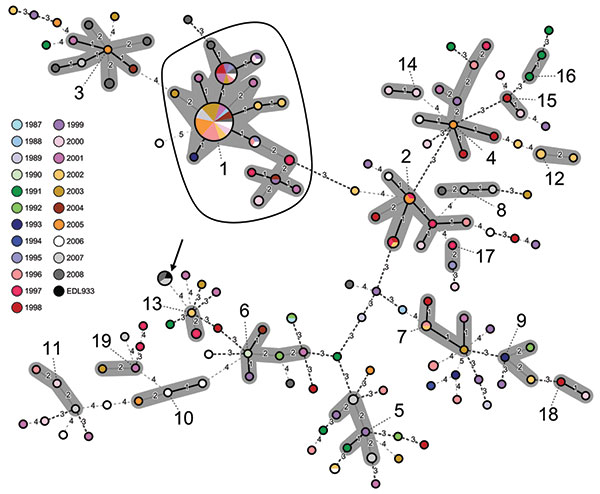
Figure. Minimum spanning tree based on multilocus variable number tandem repeat analysis (MLVA) allelic profiles, showing the phylogenetic relationship between 202 epidemiologically unrelated enterohemorrhagic Escherichia coli (EHEC) O157:H7/H– strains, Germany, 1987–2008. Each node represents a unique MLVA profile. Size of the nodes is proportional to the number of isolates per MLVA profile. Small numbers and specific dashed lines between nodes represent distance between the nodes, i.e., number of different alleles. Clusters are indicated by numbers according to their size (Table 2) and are highlighted in gray. All strains within the enclosed area are sorbitol-fermenting EHEC O157:H–. The arrangement of colors within each node reflects the proportion of strains isolated from different years per node and corresponding MLVA profile. Arrow indicates reference strain E. coli O157:H7 EDL933.