Volume 18, Number 7—July 2012
Dispatch
Role of Birds in Dispersal of Etiologic Agents of Tick-borne Zoonoses, Spain, 2009
Figure
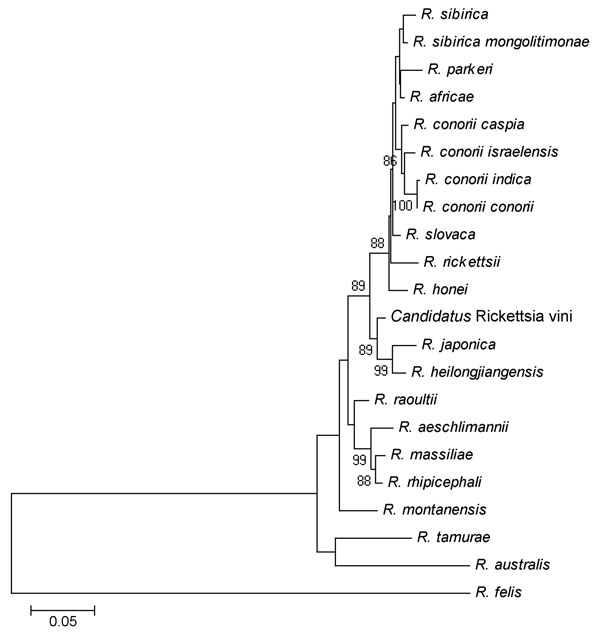
Figure. . . The phylogenetic position of Candidatus Rickettsia vini based on the ompA nucleotide sequences in a study of the role of birds in dispersal of etiologic agents of tick-borne zoonoses, Spain, 2009. The evolutionary history was inferred by using the neighbor-joining method. The optimal tree with the sum of branch length = 1.09961140 is shown. The percentage of replicate trees in which the associated taxa clustered in the bootstrap test (1,000 replicates) is shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed by using the Kimura 2-parameter method and are in the units of the number of base substitutions per site. Codon positions included were 1st+2nd+3rd+Noncoding. All positions containing gaps and missing data were eliminated from the dataset. A total of 563 positions were in the final dataset. Phylogenetic analyses were conducted in MEGA4 (16 in Technical Appendix).