Volume 19, Number 6—June 2013
Research
Novel Mycobacterium tuberculosis Complex Isolate from a Wild Chimpanzee
Figure 1
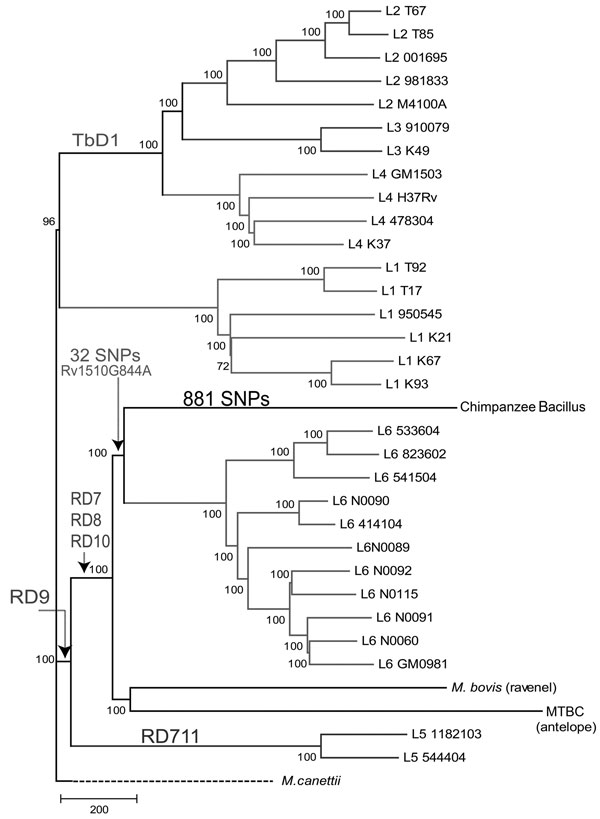
Figure 1. . Neighbor-joining phylogenic tree constructed on the basis of 13,480 variable common nucleotide positions across 36 human and animal Mycobacterium tuberculosis complex (MTBC) genome sequences, including 21 previously published genomes (18) and the MTBC strain isolated from an adult female chimpanzee that was found dead in Taï National Park, Côte d’Ivoire, on August 5, 2009 (Chimpanzee Bacillus). The tree is rooted with M. canettii, the closest known outgroup. Node support after 1,000 bootstrap replications is indicated. Genomic deletions identified in (7) are indicated. The number of single-nucleotide polymorphisms (SNPs) exclusive of the chimpanzee strain is indicated in the respective branch, and the number of SNPs shared with the most closely related group of strains is indicated in the common branch. Scale bar indicates number of SNPs. This tree is congruent with the maximum-likelihood phylogeny shown in Technical Appendix Figure 2.
References
- Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–44 . DOIPubMedGoogle Scholar
- de Jong BC, Antonio M, Gagneux S. Mycobacterium africanum—review of an important cause of human tuberculosis in West Africa. PLoS Negl Trop Dis. 2010;4:e744 . DOIPubMedGoogle Scholar
- Gutierrez MC, Brisse S, Brosch R, Fabre M, Omaïs B, Marmiesse M, Ancient origin and gene mosaicism of the progenitor of Mycobacterium tuberculosis. PLoS Pathog. 2005;1:e5 . DOIPubMedGoogle Scholar
- Brosch R, Gordon SV, Marmiesse M, Brodin P, Buchrieser C, Eiglmeier K, A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc Natl Acad Sci U S A. 2002;99:3684–9. DOIPubMedGoogle Scholar
- Huard RC, Fabre M, de Haas P, Claudio Oliveira Lazzarini L, van Soolingen D, Cousins D, Novel genetic polymorphisms that further delineate the phylogeny of the Mycobacterium tuberculosis. J Bacteriol. 2006;188:4271–87. DOIPubMedGoogle Scholar
- van Ingen J, Rahim Z, Mulder A, Boeree MJ, Simeone R, Brosch R, Characterization of Mycobacterium orygis as M. tuberculosis complex subspecies. Emerg Infect Dis. 2012;18:653–5 . DOIPubMedGoogle Scholar
- Gagneux S, DeRiemer K, Van T, Kato-Maeda M, de Jong BC, Narayanan S, Variable host-pathogen compatibility in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A. 2006;103:2869–73. DOIPubMedGoogle Scholar
- Mostowy S, Cousins D, Brinkman J, Aranaz A, Behr M. Genomic deletions suggest a phylogeny for the Mycobacterium tuberculosis complex. J Infect Dis. 2002;186:74–80. DOIPubMedGoogle Scholar
- Garnier T, Eiglmeier K, Camus JC, Medina N, Mansoor H, Pryor M, The complete genome sequence of Mycobacterium bovis. Proc Natl Acad Sci U S A. 2003;100:7877–82. DOIPubMedGoogle Scholar
- Smith NH, Kremer K, Inwald J, Dale J, Driscoll JR, Gordon SV, Ecotypes of the Mycobacterium tuberculosis complex. J Theor Biol. 2006;239:220–5. DOIPubMedGoogle Scholar
- Cousins DV, Peet RL, Gaynor WT, Williams SN, Gow BL. Tuberculosis in imported hyrax (Procavia capensis) caused by an unusual variant belonging to the Mycobacterium tuberculosis complex. Vet Microbiol. 1994;42:135–45. DOIPubMedGoogle Scholar
- Alexander KA, Laver PN, Michel AL, Williams M, van Helden PD, Warren RM, Novel Mycobacterium tuberculosis complex pathogen, M. mungi. Emerg Infect Dis. 2010;16:1296–9. DOIPubMedGoogle Scholar
- Leendertz FH, Pauli G, Maetz-Rensing K, Boardman W, Nunn C, Ellerbrok H, Pathogens as drivers of population declines: the importance of systematic monitoring in great apes and other threatened mammals. Biol Conserv. 2006;131:325–37. DOIGoogle Scholar
- Reddington K, O’Grady J, Dorai-Raj S, Niemann S, van Soolingen D, Barry T. A novel multiplex real-time PCR for the identification of Mycobacteria associated with zoonotic tuberculosis. PLoS ONE. 2011;6:e23481. DOIPubMedGoogle Scholar
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Current protocols in molecular biology. New York: Greene Publishing Associated and Wiley-Interscience; 1987.
- Gagneux S, Small PM. Global phylogeography of Mycobacterium tuberculosis and implications for tuberculosis product development. Lancet Infect Dis. 2007;7:328–37. DOIPubMedGoogle Scholar
- Hershberg R, Lipatov M, Small PM, Sheffer H, Niemann S, Homolka S, High functional diversity in Mycobacterium tuberculosis driven by genetic drift and human demography. PLoS Biol. 2008;6:e311. DOIPubMedGoogle Scholar
- Li H, Durbin R. Fast and accurate short read alignment with Burrows Wheeler transform. Bioinformatics. 2009;25:1754–60. DOIPubMedGoogle Scholar
- Comas I, Chakravartti J, Small PM, Galagan J, Niemann S, Kremer K, Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat Genet. 2010;42:498–503. DOIPubMedGoogle Scholar
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9 . DOIPubMedGoogle Scholar
- Chevreux B, Wetter T, Suhai S. Genome sequence assembly using trace signals and additional sequence information. In: Computer science and biology: proceedings of the German Conference on Bioinformatics. Braunschweig (Germany): GBF Braunschweig Department of Bioinformatics; 1999. p. 45–56.
- Rissman AI, Mau B, Biehl BS, Darling AE, Glasner JD, Perna NT. Reordering contigs of draft genomes using the Mauve Aligner. Bioinformatics. 2009;25:2071–3. DOIPubMedGoogle Scholar
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–9. DOIPubMedGoogle Scholar
- Posada D. jModelTest: Phylogenetic model averaging. Mol Biol Evol. 2008;25:1253–6. DOIPubMedGoogle Scholar
- Guindon S, Lethiec F, Duroux P, Gascuel O. PHYML Online—a web server for fast maximum likelihood-based phylogenetic inference. Nucleic Acids Res. 2005;33:W557–9. DOIPubMedGoogle Scholar
- Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19:1572–4. DOIPubMedGoogle Scholar
- Kamerbeek J, Schouls L, Kolk A, van Agterveld M, van Soolingen D, Kuijper S, Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J Clin Microbiol. 1997;35:907–14 .PubMedGoogle Scholar
- Goh KS, Fabre M, Huard RC, Schmid S, Sola C, Rastogi N. Study of the gyrB gene polymorphism as a tool to differentiate among Mycobacterium tuberculosis complex subspecies further underlines the older evolutionary age of Mycobacterium canettii. Mol Cell Probes. 2006;20:182–90. DOIPubMedGoogle Scholar
- Bentley SD, Comas IÃ, Bryant JM, Walker D, Smith NH, Harris SR, The genome of Mycobacterium africanum West African 2 reveals a lineage-specific locus and genome erosion common to the M. tuberculosis complex. PLoS Negl Trop Dis. 2012;6:e1552.
- Demay C, Liens B, Burguiere T, Hill V, Couvin D, Millet J, SITVITWEB—a publicly available international multimarker database for studying Mycobacterium tuberculosis genetic diversity and molecular epidemiology. Infect Genet Evol. 2012;12:755–66. DOIPubMedGoogle Scholar
- Mostowy S, Onipede A, Gagneux S, Niemann S, Kremer K, Desmond EP, Genomic analysis distinguishes Mycobacterium africanum. J Clin Microbiol. 2004;42:3594–9. DOIPubMedGoogle Scholar
- Thorel MF. Isolation of Mycobacterium africanum from monkeys. Tubercle. 1980;61:101–4 . DOIPubMedGoogle Scholar
- Michel AL, Venter L, Espie IW, Coetzee ML. Mycobacterium tuberculosis infections in eight species at the national zoological gardens of South Africa, 1991−2001. J Zoo Wildl Med. 2003;34:364–70. DOIPubMedGoogle Scholar
- Mostowy S, Cousins D, Behr MA. Genomic interrogation of the Dassie Bacillus reveals it as a unique RD1 mutant within the Mycobacterium tuberculosis complex. J Bacteriol. 2004;186:104–9 . DOIPubMedGoogle Scholar
- Wolfe ND, Dunavan CP, Diamond J. Origins of major human infectious diseases. Nature. 2007;447:279–83. DOIPubMedGoogle Scholar
- Calvignac-Spencer S, Leendertz SAJ, Gillespie TR, Leendertz FH. Wild great apes as sentinels and sources of infectious disease. Clin Microbiol Infect. 2012;18:521–7. DOIPubMedGoogle Scholar
- Boesch C, Boesch-Achermann H. The chimpanzees of the Taý Forest: behavioural ecology and evolution. Oxford/New York: Oxford University Press; 2000.
- Koeck JL, Fabre M, Simon F, Daffe M, Garnotel E, Matan AB, Clinical characteristics of the smooth tubercle bacilli Mycobacterium canettii infection suggest the existence of an environmental reservoir. Clin Microbiol Infect. 2011;17:1013–9. DOIPubMedGoogle Scholar
1These authors contributed equally to this article.