Volume 21, Number 7—July 2015
Dispatch
Gastroenteritis Outbreaks Caused by Norovirus GII.17, Guangdong Province, China, 2014–2015
Figure 2
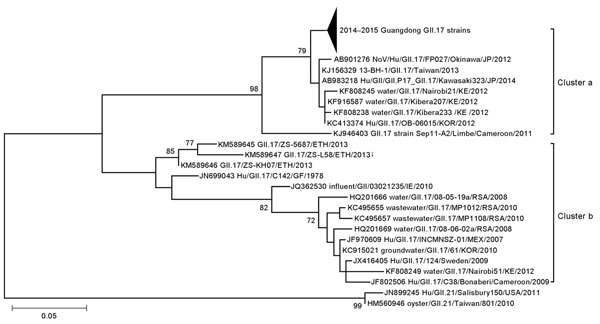
Figure 2. Phylogenetic tree of noroviruses based on the 282-bp region of the capsid N terminus/shell gene. Nucleotide sequences were analyzed by using the maximum-likelihood method. Supporting bootstrap values >70 are shown. The subtrees of GII.17 detected in Guangdong Province, China, during 2014–2015 were compressed. GII.21 genotype strains were used as outgroups. Scale bar indicates nucleotide substitutions per site. Sequences of 24 reference norovirus strains are included. Arrowhead represents number of strains from Guangdong, 2014–2015. ETH, Ethiopia; GF, French Guiana; IE, Ireland; JP, Japan; KE, Kenya; KOR, Korea; RSA, The Republic of South Africa; MEX, Mexico.
1These authors contributed equally to this article.
Page created: June 16, 2015
Page updated: June 16, 2015
Page reviewed: June 16, 2015
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.