Volume 22, Number 3—March 2016
Letter
Novel Reassortant Avian Influenza A(H5N1) Virus in Human, Southern Vietnam, 2014
To the Editor: The first case of human infection with highly pathogenic avian influenza A(H5N1) virus in Vietnam was reported in December 2003 (1), and >120 human cases were confirmed through 2013, with a high case-fatality rate (2). In 2013, clade 2.3.2.1a/c H5N1 viruses circulated widely in poultry across the country, although clade 1.1.1/1.1.2 H5N1 viruses predominated in poultry from the Mekong Delta region to central Vietnam (3,4).
In 2014, two cases of human infection with A(H5N1) virus were identified in southern Vietnam. One case was associated with a clade 1.1.2 reassortant virus, A/Vietnam/14012902/2014 (Global Initiative on Sharing All Influenza Data [GISAID; http://www.gisaid.org] accession nos. EPI624919–EPI624926), which had been previously detected in Cambodia and Vietnam (5,6). We isolated the virus from the other case, performed phylogenetic analysis to identify the clade of this virus, and identified a novel virus that had undergone gene reassortment.
The case-patient was a 52-year-old man who lived in Binh Phuoc Province (140 km northeast of Ho Chi Minh City). On January 11, 2014, he experienced mild fever and general fatigue; high fever developed on January 13. He was hospitalized with dyspnea on January 16 and died 2 days later. He was not given antiviral drug treatment. Dead poultry infected with H5N1 viruses were found scattered near his house during January 1–16, and he buried his 2 dead chickens on January 5. H5N1 virus infection was detected in the patient’s throat swab specimen by real-time reverse transcription PCR at the Pasteur Institute in Ho Chi Minh City. Virus was isolated by inoculating the throat swab specimens into 10-day-old embryonated chicken eggs; the resulting isolate, A/Vietnam/14011801/2014 (GISAID accession nos. EPI624911–EPI624918), then underwent gene sequencing. The 8 viral genes were amplified with SuperScript III Reverse Transcriptase Kit (Fisher Scientific, Pittsburg, PA, USA) and Phusion High-Fidelity DNA Polymerase (New England BioLabs, Ipswich, MA, USA) with specific paired primers, according to the manufacturer’s instructions, and sequenced on an ABI 3730 automated sequencer with Big-Dye Terminator Cycle Sequencing reagents (Applied Biosystems, Foster City, CA, USA). Whole genome sequence was determined.
By gene sequencing analysis, A/Vietnam/14011801/2014 was found to have the multibasic cleavage site of hemagglutinin (HA) protein, which indicates highly pathogenic avian influenza A(H5N1) viruses, and was shown to predict binding specificity to an avian α2,3 sialic acid receptor. The neuraminidase gene possessed no amino acid substitutions associated with decreased antiviral activity, nor did the virus have amino acid substitutions associated with increased adaptation, virulence, infectivity, or transmissibility in mammalian hosts, including the E627K and D701N mutations in polymerase basic protein 2 (7).
Phylogenetic analyses of the 8 viral genes of A/Vietnam/14011801/2014 were performed by using databases (GISAID and the Influenza Virus Resource, National Center for Biotechnology Information, Bethesda, MD, USA; http://www.ncbi.nlm.nih.gov/genomes/FLU/FLU.html) that contained complete sequences of viral genomes belonging to clades 1.1.1, 1.1.2, and 2.3.2.1 a/b/c, most of which were collected in Vietnam, particularly after 2012. Neighbor-joining and Kimura 2-parameter methods were implemented by using MEGA version 5.0 software (http://www.megasoftware.net). Reliability of the phylogenetic analysis was tested by using 1,000 bootstrap replications. Lineages of the HA gene were defined by using previously described criteria (8). Lineages of the other 7 genes were defined by using criteria and nomenclature of Nguyen et al. (9).
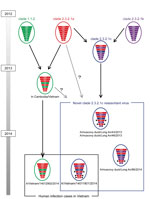
The HA of A/Vietnam/14011801/2014 belonged to clade 2.3.2.1c (online Technical Appendix Figure, panel A, http://wwwnc.cdc.gov/EID/article/23/3/15-1360-Techapp.pdf). The neuraminidase, polymerase basic proteins 1 and 2, and polymerase acid protein genes of this virus were also derived from respective lineages of ancestor clade 2.3.2.1c (Technical Appendix Figure, panels B–E). However, nucleoprotein, matrix, and nonstructural genes were classified as lineages of ancestor clade 2.3.2.1a (online Technical Appendix Figure, panels F–H) and differed from the gene lineages of almost all clade 2.3.2.1c viruses isolated from poultry in Vietnam. As reported in the Influenza Virus Resource, 2 viruses collected in Vietnam in December 2013 (A/muscovy duck/Long An/43/2013 and A/muscovy duck/Long An/46/2013) were similar reassortant viruses of clade 2.3.2.1c (Figure). However, the ancestor of the nonstructural gene lineage of these 2 viruses is clade 2.3.2.1c, which differs from A/Vietnam/14011801/2014. The differences indicate that A/Vietnam/14011801/2014 is a novel reassortant virus between clades 2.3.2.1a and 2.3.2.1c, between clades 1.1.2 and 2.3.2.1c, or both (Figure). This novel reassortant virus has not been reported in poultry in Vietnam, although novel reassortants between clade 1.1.2 and clade 2.3.2.1a viruses have been detected in Vietnam since 2013 (i.e., A/Vietnam/VP13-28H/2013, GISAID accession nos. EPI624927–EPI624934; and A/Vietnam/14012902/2014) (6). These novel reassortment viruses were first identified in human, animal, and environmental samples in Cambodia in 2013 (5). Other novel gene reassortments in clade 2.3.2.1 viruses have been previously reported (10), and new clade 2.3.4.4 viruses have been observed in Vietnam since 2014.
As multiple clade viruses co-circulate, reassortment events occur frequently in Vietnam. Continuous surveillance of avian influenza A(H5N1) viruses, not only in humans but also in poultry and wild birds, is needed for infection control measures during epidemics of these viruses.
Acknowledgment
This study was partially supported by Grants-in-Aid for Emerging and Reemerging Infectious Diseases from the Ministry of Health, Labour and Welfare, Japan.
References
- Tran TH, Nguyen TL, Nguyen TD, Luong TS, Pham PM, Nguyen V, et al. Avian influenza A (H5N1) in 10 patients in Vietnam. N Engl J Med. 2004;350:1179–88. DOIPubMedGoogle Scholar
- World Health Organization. Cumulative number of confirmed human cases for avian influenza A(H5N1) reported to WHO, 2003–2014 [cited 2015 Jan 6]. http://www.who.int/influenza/human_animal_interface/EN_GIP_20141002CumulativeNumberH5N1cases.pdf?ua=1
- Nguyen DT, Bryant JE, Davis CT, Nguyen LV, Pham LT, Loth L, Prevalence and distribution of avian influenza a(H5N1) virus clade variants in live bird markets of Vietnam, 2011–2013. Avian Dis. 2014;58:599–608. DOIPubMedGoogle Scholar
- Lee EK, Kang HM, Kim KI, Choi JG, To TL, Nguyen TD, Genetic evolution of H5 highly pathogenic avian influenza virus in domestic poultry in Vietnam between 2011 and 2013. Poult Sci. 2015;94:650–61. DOIPubMedGoogle Scholar
- Rith S, Davis CT, Duong V, Sar B, Horm SV, Chin S, Identification of molecular markers associated with alteration of receptor-binding specificity in a novel genotype of highly pathogenic avian influenza A(H5N1) viruses detected in Cambodia in 2013. J Virol. 2014;88:13897–909. DOIPubMedGoogle Scholar
- Thor SW, Nguyen H, Balish A, Hoang AN, Gustin KM, Nhung PT, Detection and characterization of clade 1 reassortant H5N1 viruses isolated from human cases in Vietnam during 2013. PLoS ONE. 2015;10:e0133867. DOIPubMedGoogle Scholar
- Le QM, Sakai-Tagawa Y, Ozawa M, Ito M, Kawaoka Y. Selection of H5N1 influenza virus PB2 during replication in humans. J Virol. 2009;83:5278–81. DOIPubMedGoogle Scholar
- World Health Organization/World Organization for Animal Health/Food and Agriculture Organization H5N1 Evolution Working Group. Revised and updated nomenclature for highly pathogenic avian influenza A (H5N1) viruses. Influenza Other Respi Viruses. 2014;8:384–8. DOIGoogle Scholar
- Nguyen T, Rivailler P, Davis CT, Hoa do T, Balish A, Dang NH, Evolution of highly pathogenic avian influenza (H5N1) virus populations in Vietnam between 2007 and 2010. Virology. 2012;432:405–16. DOIPubMedGoogle Scholar
- Creanga A, Thi Nguyen D, Gerloff N, Thi Do H, Balish A, Dang Nguyen H, Emergence of multiple clade 2.3.2.1 influenza A (H5N1) virus subgroups in Vietnam and detection of novel reassortants. Virology. 2013;444:12–20.DOIPubMedGoogle Scholar
Figure
Cite This Article1These first authors contributed equally to this article.
Related Links
Table of Contents – Volume 22, Number 3—March 2016
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Tsutomu Kageyama, Influenza Virus Research Center, National Institute of Infectious Diseases, 4-7-1 Gakuen, Musashimurayama-shi, Tokyo 208-0011, Japan
Top