Volume 25, Number 10—October 2019
Dispatch
Two Cases of Borrelia miyamotoi Meningitis, Sweden, 2018
Figure 2
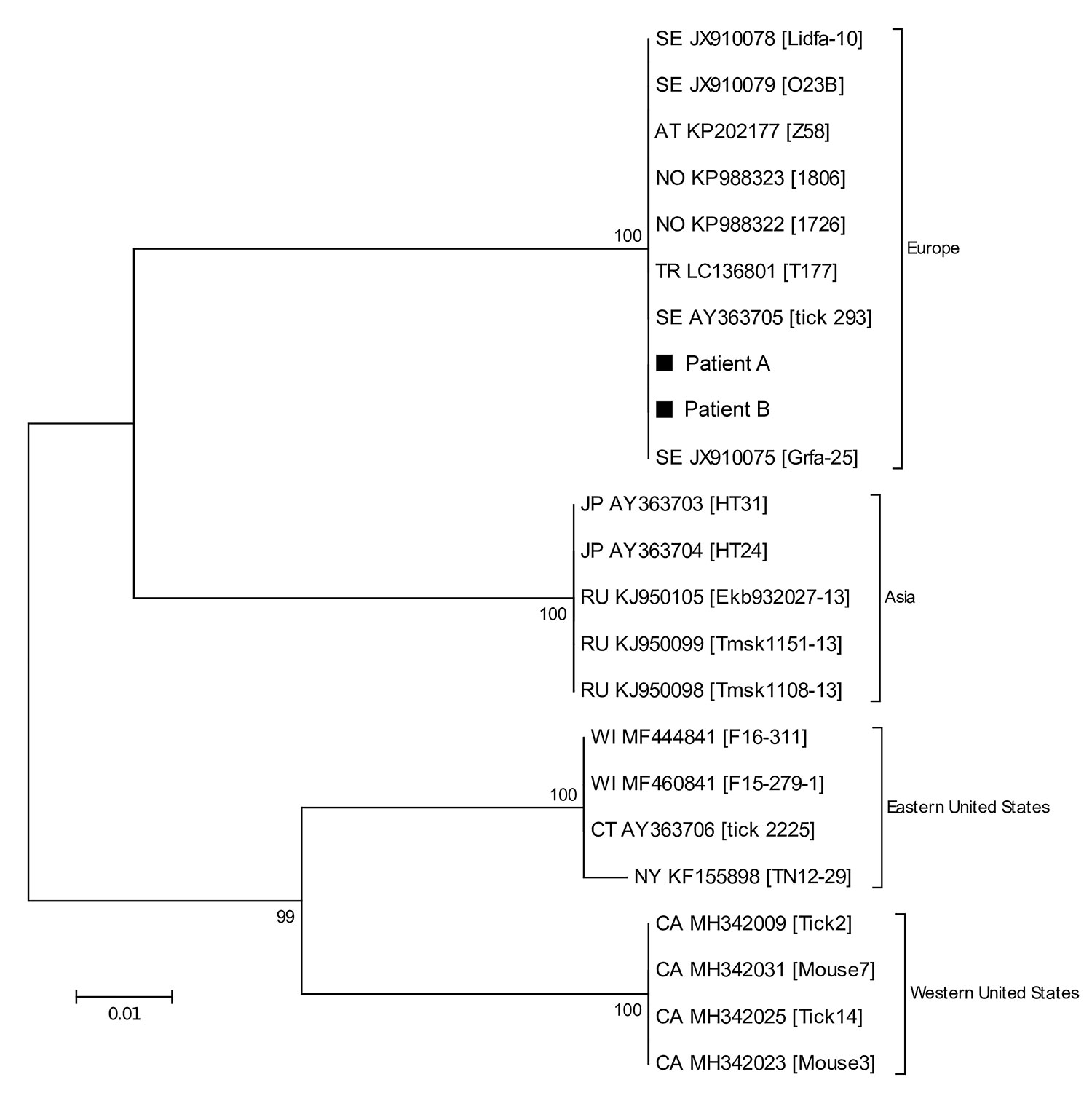
Figure 2. Phylogenetic tree based on 16S-23S intergenic spacer region sequences of Borrelia miyamotoi from 2 patients in Sweden, 2018 (patients A and B, black squares), and reference sequences. Tree constructed using the maximum-likelihood method based on the Tamura-Nei model and complete deletion. Sequences detected from patients in this study were deposited into GenBank under accession nos. MK458687 (patient A) and MK458688 (patient B). The source of each reference sequence is indicated by an accession number preceded by a state or country code: AT, Austria; CA, California; CT, Connecticut; JP, Japan; NO, Norway; NY, New York; RU, Russian Federation; SE, Sweden; TR, Turkey; WI, Wisconsin. The accession number is followed by the isolate name in brackets. The reliability of the tree was tested by 500 bootstrap replicate analyses; only values >50% are shown. The phylogenetic relationship between the B. miyamotoi strains detected in our patients was corroborated by the DNA sequences obtained from the glpQ and p66 genes (data not shown). Scale bar indicates nucleotide substitutions per site.