Volume 26, Number 3—March 2020
Research
Whole-Genome Sequencing to Detect Numerous Campylobacter jejuni Outbreaks and Match Patient Isolates to Sources, Denmark, 2015–2017
Figure 3
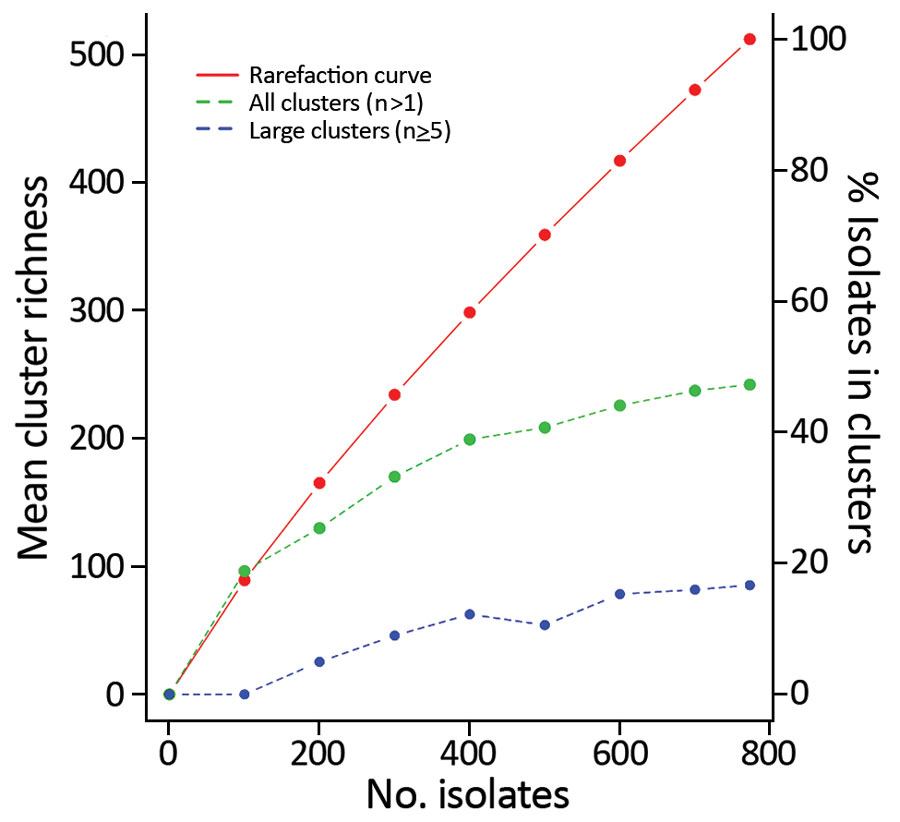
Figure 3. Rarefaction curve of Campylobacter jejuni clinical isolates from Denmark, 2015–2017. Curve of cluster types is shown relating to the left y-axis, revealing 512 different cluster types, which encompass the 104 defined clusters as well as the 408 sporadic cases assigned to their own cluster type. The right y-axis indicates the percentage of clinical isolates assigned to any cluster or a large cluster for a given subsample size. The rarefaction curve indicates that only a small fraction of the diversity is sampled; a larger fraction of the sporadic isolates would be included in clusters if a greater fraction of the population had been sampled.
Page created: February 20, 2020
Page updated: February 20, 2020
Page reviewed: February 20, 2020
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.