Novel SARS-CoV-2 Variant Derived from Clade 19B, France
Slim Fourati
, Jean-Winoc Decousser, Souraya Khouider, Melissa N’Debi, Vanessa Demontant, Elisabeth Trawinski, Aurélie Gourgeon, Christine Gangloff, Grégory Destras, Antonin Bal, Laurence Josset, Alexandre Soulier, Yannick Costa, Guillaume Gricourt, Bruno Lina, Raphaël Lepeule, Jean-Michel Pawlotsky, and Christophe Rodriguez
Author affiliations: Institut Mondor de Recherche Biomédicale, Université Paris-Est, Créteil, France; (S. Fourati, S. Khouider, A. Gourgeon, A. Soulier, J.-M. Pawlotsky, C. Rodriguez); Hôpital Henri Mondor, Créteil (J.-W. Decousser, M. N’Debi, V. Demontant, E. Trawinski, G. Gricourt, R. Lepeule, C. Rodriguez); Hôpital Albert Chenevier, Créteil (C. Gangloff); Université de Lyon, France (G. Destras, A. Bal, L. Josset, B. Lina); Grand Hôpital de l’Est Francilien, Jossigny, France (Y. Costa)
Main Article
Figure
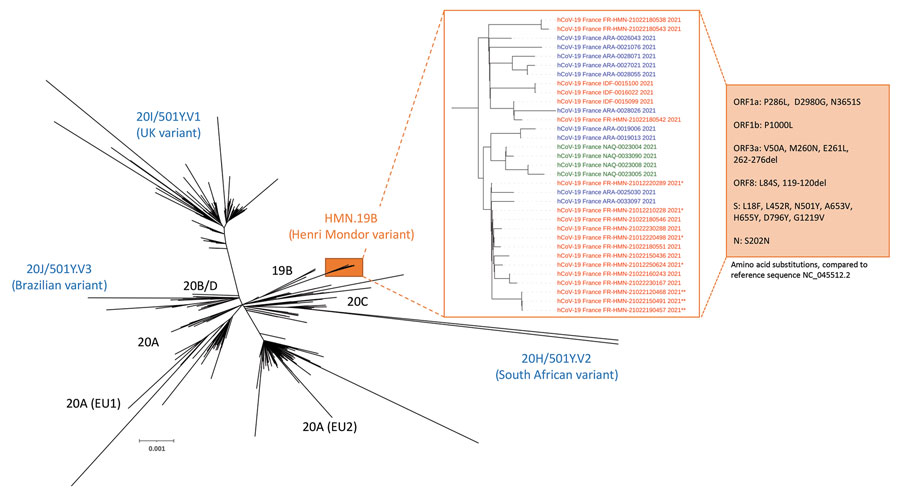
Figure. Phylogenetic tree built with sequences from the 33 patients infected with the new HMN.19B or Henri Mondor variant and from 1,537 SARS-CoV-2–infected patients in France sampled during January 18–February 23, 2021, sequenced in 9 successive series. Phylogeny was performed after full-length genome alignment with Muscle 3.8.31 (maximum-likelihood model general time-reversible plus invariant sites model, 1,000 bootstrap replicates) by means of IQ-Tree 1.3.11.1 and iTOL. The HMN.19B (Henri Mondor) variant cluster is considerably different from all the others, with a 99% bootstrap value. HMN.19B sequences are colored according to the geographic origin of the patients: red, greater Paris area (IDF or HMN); blue, southeast France (ARA); green, southwest France (NAQ). *Original cluster in an occupational health unit; **cluster in the hematology department of our institution. Amino-acid substitutions and deletions detected in all sequences are described in the orange box. ARA, Auvergne-Rhône-Alpes; HMN, Henri Mondor; IDF, Ile-de-France; NAQ, Nouvelle-Aquitaine.
Main Article
Page created: March 16, 2021
Page updated: April 22, 2021
Page reviewed: April 22, 2021
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
