Volume 5, Number 6—December 1999
Dispatch
New Rickettsiae in Ticks Collected in Territories of the Former Soviet Union
Abstract
Dermacentor nuttallii from Siberia, Rhipicephalus sanguineus from Crimea, and Rh. pumilio from the Astrakhan region were infected with Rickettsia sibirica (12%), R. conorii (8%), and the Astrakhan fever agent (3%), respectively. Three new Rickettsiae of the R. massiliae genogroup were identified in ticks by 16S rDNA, gltA, and ompA sequencing.
Rickettsiae are obligate intracellular gram-negative bacteria associated with arthropod vectors, ticks, mites, and insects that, while feeding, can transmit Rickettsiae to animals and humans. The rickettsioses have characteristic clinical features, including fever, headache, maculopapular eruption, and sometimes eschar formation (primary lesion). The number of representatives of the genus Rickettsia has increased over the last decades as a result of improved cell culture isolation and agent identification techniques (1). Sequence comparison of gene coding for citrate synthase (gltA) (2), rOmpA outer membrane protein (ompA) (3), and 16S rRNA (4) has become the most reliable method of identifying Rickettsiae (5-8). We describe polymerase chain reaction (PCR) amplification and sequence determination to identify Rickettsiae in naturally infected ixodid ticks in three regions of Russia endemic for tickborne rickettsioses.
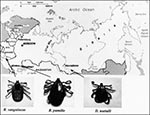
Rhipicephalus pumilio ticks (65 adults) were collected in 1996 from dogs in the Astrakhan region. Dermacentor nuttallii ticks (101 adults) were collected in 1994 in the village of Verhnyi Kouus, the Altay Mountains, Siberia. In 1997, Rh. sanguineus ticks (2 adults and 35 nymphs) were collected in the town of Saki, Crimea region, from dogs whose owners had serologic evidence of Mediterranean spotted fever (Figure 1). The ticks were kept at room temperature before being washed in iodized alcohol (10 minutes) just before testing, rinsed in distilled water, and dried on sterile filter paper. DNA was extracted from ticks by using the QIAmp Tissue Kit (QIAGEN, Hilden, Germany). Rickettsial DNA was detected by PCR with primers specific for Rickettsiae: RpCS.877p-RpCS.1273r, which amplify a 396-bp fragment of gltA (2), and Rr190.70p-190-701 (3), which amplify a fragment of ompA from 629 to 632 bp. For all positive ticks, 587 to 590 bp of ompA were sequenced by using the ABI PRISM Dye Terminator Cycle Sequencing Kit with Amplitap Polymerase FS (PE Applied Biosystems, Warrington WA1 4SR, UK). Sequences were analyzed with the Applied Biosystem 377 automatic sequencing system. For newly detected genotypes, sequences of 16S rRNA encoding gene, gltA, and ompA were determined as previously described (2-4) (see Figure 2 for GenBank codes).
We detected two different Rickettsiae in Rh. pumilio (Astrakhan fever agent and RpA4 genotype); two Rickettisiae from D. nutallii in Siberia (Rickettsia sibirica and DnS14 and DnS 28 genotypes); and R. conorii from Rh. sanguineous ticks in Crimea (Table).
Our results confirm previous data of high epidemic activity of the Altay focus for North Asian tick typhus and the crucial role of D. nuttallii as a reservoir of R. sibirica infection (9). Our results are also consistent with those of a study in 1991 based on hemolymph testing (10), in which 3.2% of ticks from the Astrakhan region were demonstrated to be infected with Rickettsiae.
An outbreak of Mediterranean spotted fever due to infection with R. conorii occurred in Crimea from 1947 to 1957. Only sporadic cases of the disease were reported (11) until 1995, when the incidence of Mediterranean spotted fever increased in central Crimea, with 40 cases in 1996 and more than 70 in 1997. Most cases occurred in the summer, when the Rh. sanguineus nymphs (principal vectors of R. conorii) (1) were active. Our results, showing that 8% of the Rh. sanguineus studied contained R. conorii DNA, provide further evidence of the Mediterranean spotted fever outbreak in the region. To date, only the R. conorii strain M-1, isolated in the territories of the former Soviet Union (the Black Sea coast of Georgia), has been genetically characterized. This strain is genetically distinct from the other strains of R. conorii, i.e., Indian tick typhus and the Moroccan and Malish strains (3). Our detection of the R. conorii strain identical to the Malish strain is the first evidence of the genetic heterogeneity of R. conorii in the region.
The ompA sequences obtained from PCR-amplified products were different from those described for the known Rickettsiae for one DNA sample extracted from Rh. pumilio from the Astrakhan region (RpA4) and four DNA samples from D. nuttallii collected in Siberia (DnS14, DnS28, DnS79, DnS94). The sequences for the samples from D. nuttalii (DnS28, DnS79, and DnS94) were identical but differed from those of DnS14 and Rh. pumilio RpA4/2.
The three new rickettsial agents were closely related and branched with members of the R. massiliae group, together with R. rhipicephali, Bar 29, R. aeschlimannii, and R. montanensis (Figure 2). Comparison of the sequences obtained by using the program BLAST demonstrated that they also differed from those of the Cadiz agent characterized from Ixodes ricinus in Spain (6), those of the Cooleyi genotype characterized from I. scapularis (5), MOA and WB-8-2 isolated from Amblyomma americanum and I. scapularis, respectively (8), and R. peacockii (7) in the United States.
The pathogenicity of the members of the R. massiliae group is unknown, and their main reservoirs are regarded as ticks of the genus Rhipicephalus for R. massiliae and Bar 29. R. aeschlimannii has been isolated from Hyalomma marginatum and R. montanensis from ticks of the genus Dermacentor. R. rhipicephali has been demonstrated in ticks of the genus Dermacentor and in Rh. sanguineus (1). The similarity of gltA, ompA, and 16S rRNA gene sequences indicates that these three new agents are close to each other (from 99.7% to 99.9%) and could constitute a new rickettsial species.
In the United States, various tickborne Rickettsiae occur in areas endemic for R. rickettsii, the agent of Rocky Mountain spotted fever (7). Similarly, in Mediterranean countries, several recently described Rickettsiae have been found in ticks of the Rh. sanguineus complex in the regions endemic for Mediterranean spotted fever caused by R. conorii (6). The effects of the presence of different Rickettsiae on the prevalence of infection rates of ticks with individual Rickettsiae and on the epidemiology of infections in humans have yet to be determined. R. sibirica and the Astrakhan fever agent are prevalent in Siberia and the Astrakhan region, respectively, but the pathogenicity of the new rickettsial genotypes has yet to be investigated.
Dr. Rydkina is a senior researcher at the Gamalaya Institute in Moscow. She was trained as a postdoctoral fellow in Marseille, France. Her main interest is in rickettsial diseases.
Acknowledgment
We thank P.J. Kelly and E. Birtles for reviewing the article.
References
- Raoult D, Roux V. Rickettsioses as paradigms of new or emerging infectious diseases. Clin Microbiol Rev. 1997;10:694–719.PubMedGoogle Scholar
- Roux V, Rydkina E, Eremeeva M, Raoult D. Citrate synthase gene comparison, a new tool for phylogenetic analysis, and its application for the Rickettsiae. Int J Syst Bacteriol. 1997;47:252–61. DOIPubMedGoogle Scholar
- Fournier P-E, Roux V, Raoult D. Phylogenetic analysis of spotted fever group rickettsiae by study of the outer surface protein rOmpA. Int J Syst Bacteriol. 1998;48:839–49. DOIPubMedGoogle Scholar
- Roux V, Raoult D. Phylogenetic analysis of the genus Rickettsia by 16S rDNA sequencing. Res Microbiol. 1995;146:385–96. DOIPubMedGoogle Scholar
- Billings AN, Teltow GJ, Weaver SC, Walker DH. Molecular characterization of a novel Rickettsia species from Ixodes scapularis in Texas. Emerg Infect Dis. 1998;4:305–9. DOIPubMedGoogle Scholar
- Marquez FJ, Muniain MA, Soriguer RC, Izquierdo G, Rodriguez-Baño J, Borobio M. Genotypic identification of an undescribed spotted fever group Rickettsia in Ixodes ricinus from southwestern Spain. Am J Trop Med Hyg. 1998;58:570–7.PubMedGoogle Scholar
- Niebylski ML, Shrumpf ME, Burgdorfer W, Ficher ER, Gage KL, Schwan TG. Rickettsia peacockii sp. nov., a new species infecting wood ticks, Dermacentor andersoni, in Western Montana. Int J Syst Bacteriol. 1997;47:446–52. DOIPubMedGoogle Scholar
- Weller SJ, Baldridge GD, Munderloh UG, Noda H, Simser J, Kurtti T. Phylogenetic placement of Rickettsiae from ticks Amblyomma americanum and Ixodes scapularis. J Clin Microbiol. 1998;36:1305–17.PubMedGoogle Scholar
- Rudakov NV. Tick-borne rickettsiosis in Russia (epidemiology and current conditions of natural foci). In: Kazar J, Toman R, editors. Rickettsiae and rickettsial diseases. Proceedings of the 5th International Symposium; 1996; Veda, Bratislava. p. 216-20.
- Eremeeva ME, Beati L, Makarova VA, Fetisova NF, Tarasevich IV, Balayeva NM, Astrakhan fever rickettsiae: antigenic and genotypic analysis of isolates obtained from human and Rhipicephalus pumilio ticks. Am J Trop Med Hyg. 1994;51:697–706.PubMedGoogle Scholar
- Kulagin SM, Tarasevich IV, Rubakin PE, Nikitin AM, Krupina ZN. On eradication of Marseilles fever [in Russian]. J Microbiol Epidemiol Immunobiol. 1960;8:117–21.
Figures
Table
Cite This Article1The received sequences of the new rickettsial agents have been deposited in GenBank. Sequences of ompA were deposited as two parts under accession numbers: DnS28, AF120018 and AF120019; DnS14, AF120021, and AF120020; RpA4, AF120022, and AF120023. The 16S rRNA gene and gltA sequences have been deposited in GenBank under accession numbers DnS28, gltA - AF120027, 16S rRNA gene AF120024; DnS14, gltA AF120028, 16S rRNA gene AF120025; RpA4, gltA - AF120029, 16S rRNA gene AF120026.
Table of Contents – Volume 5, Number 6—December 1999
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Didier Raoult, Unité des Rickettsies, CNRS UPRES-A 6020, Faculté de Médecine, Université de la Méditerranée, 27 bd. J. Moulin 13385 Marseille, Cedex 05, France; fax: 33-49-183-0390
Top