Volume 7, Number 4—August 2001
THEME ISSUE
West Nile Virus
West Nile Virus
Rapid Determination of HLA B*07 Ligands from the West Nile Virus NY99 Genome
Abstract
Defined T cell epitopes for West Nile (WN) virus may be useful for developing subunit vaccines against WN virus infection and diagnostic reagents to detect WN virus-specific immune response. We applied a bioinformatics (EpiMatrix) approach to search the WN virus NY99 genome for HLA B*07 restricted cytotoxic T cell (CTL) epitopes. Ninety-five of 3,433 WN virus peptides scored above a predetermined cutoff, suggesting that these would be likely to bind to HLA B*07 and would also be likely candidate CTL epitopes. Compared with other methods for genome mapping, derivation of these ligands was rapid and inexpensive. Major histocompatibility complex ligands identified by this method may be used to screen T cells from WN virus-exposed persons for cell-mediated response to WN virus or to develop diagnostic reagents for immunopathogenesis studies and epidemiologic surveillance.
West Nile (WN) virus is the cause of a potentially fatal form of viral encephalitis that suddenly emerged in the New York City area during 1999. The virus is a member of the Flavivirus family, which includes St. Louis encephalitis (SLE), Japanese encephalitis (JE), hepatitis C, and dengue viruses (1,2). WN virus is common in West Asia, Africa, and the Middle East but was not reported in the Americas until the New York outbreak in 1999. The source of virus introduction to New York City is unknown; potential sources include an infected host (human or bird), an infected vector (mosquito), or bioterrorism (1,3). The WN-NY99 virus associated with the New York 1999 outbreak appears to have been circulating in Israel since 1997 (1). Other close relatives to the WN-NY99 virus were isolated in Italy (1998), Morocco (1996), Romania (1996), and Africa (1989, 1993, and 1998).
Surveillance data indicate intensified transmission and geographic expansion of the WN virus-NY99 outbreak in the northeastern United States during 1999 and 2000. Twelve states and the District of Columbia reported WN virus activity in 2000, a substantial increase over the four states reporting activity in 1999. WN-NY99 is expected to continue to spread along the east coast of the United States in 2001 and thereafter, as a result of overwintering of mosquitoes and avian migratory patterns (4,5).
WN virus's genome is 11,000 nucleotides long. The following structural proteins have been identified: envelope glycoprotein (env gp E), capsid (C), and premembrane protein (prM). The following nonstructural proteins have also been identified: NS-1, NS-2A/NS-2B, NS3, NS-4A/NS-4B; and NS-5 (RNA-directed polymerase) (1). Isolates that have been completely sequenced include the WN-NY99 virus originally obtained from a Chilean flamingo, a WN-NY99 equine isolate, the Italy 1998 virus, the Romania 1996 virus, and the prototype Eg101 virus. Although the latter viruses are closely related to WN-NY99, they are not identical to each other or to WN-NY99 (6). A virus isolated in Israel, Israel 1998, appears to be identical to WN-NY99; completion of its genome sequence is under way at the Institute Pasteur, France.
Immunology and Immunopathogenesis
An extensive body of research is available on the immunology of flaviviruses in the murine model; however, relatively little research has been done on human immune response to WN virus. Some information on human T cell responses to related viruses (e.g., JE virus, dengue) has been obtained (7,8). Both CD4 T helper cells and cytotoxic T cells that respond to JE virus and dengue proteins have been identified, and their epitopes have been mapped (9). Some of the JE virus CD4 T epitopes are identical or nearly identical to sequences in WN virus (10). Langerhans cells in the epidermis may play a role in the upregulation of immune response to the virus, processing antigen and presenting it to T cells (11,12). Mobilization of dendritic cells and antigen presentation by these cells to T cells in the lymphoid follicles may be involved in the development of immune responses to WN virus (13).
Cytotoxic T cell responses (restricted by class I major histocompatibility complex [MHC] and MHC class II) and T helper responses (restricted by class II MHC) appear to be critical components of human immune response to members of the flavivirus family (14,15). Cell-mediated immunity to WN virus may prove to be an important barrier to infection of the central nervous system, and vaccines that promote the development of T effector cells may provide protection from WN virus encephalitis or may be used to treat patients who have WN virus-related illnesses. Further research to test these hypotheses will require the development of reagents such as the T cell epitopes defined in this study.
Applying Bioinformatics to Defining T Cell Epitopes
New bioinformatics tools developed by the TB/HIV Research Lab and EpiVax (Providence, RI) enable researchers to move rapidly from genome sequence to epitope selection (16). EpiMatrix is a computer-driven pattern-matching algorithm that identifies T cell epitopes. BlastiMer permits the analysis of protein sequences for homology with other known proteins.
The goal of this project was to demonstrate the utility of a bioinformatics and computational immunology approach for the rapid selection of T cell epitope reagents. Defining these reagents will permit the evaluation of cell-mediated responses in the immunopathogenesis of WN virus, promote the development of diagnostic reagents such as tetramers (17), and provide components for epitope-based preventive or therapeutic vaccines (18-20). A secondary goal was to determine the time required to select and screen epitope candidates in vitro, since time may be a critical factor in the development of vaccines and diagnostic reagents in response to emerging infectious pathogens.
On the basis of experience with the EpiMatrix HLA B*07 prediction tool, we selected peptides for this pilot study that were expected to be restricted by HLA B*07. In studies of HIV-1 peptides, 60% of peptides selected by EpiMatrix HLA B*07 stimulated T cell responses in vitro. We therefore expected that approximately 60% of WN virus peptides selected by the same criteria would bind to HLA B*07 and stimulate T cell responses.
We screened 16 WN virus peptides and identified 12 epitope candidates, 5 of which exhibited strong binding to HLA B*07 at a range of peptide concentrations in vitro. The largest source of delay in the screening process was peptide synthesis (4 weeks from placement of order to receipt of the first set of peptides and 8 weeks until delivery of the final set of peptides). This process could be accelerated if more rapid access to MHC ligands were necessary.
The binding studies we describe are a first step to confirming immunogenicity. In cases such as WN virus, in which access to T cells from infected persons is limited, both the bioinformatics step and the binding assays can be carried out without clinical specimens. Once the epitope candidates selected by this method are confirmed in cytotoxic T cell (CTL) assays, they may be useful for 1) screening exposed persons for T cell responses, 2) investigating the immunopathogenesis of WN virus disease in humans, 3) as components of diagnostic kits developed for WN virus surveillance, 4) as reagents for measuring WN virus vaccine-related immune responses, and possibly 5) as components of a subunit vaccine for WN virus. Confirmation of T cell response to the peptides will depend on availability of peripheral blood cells from WN virus-infected patients during the 2001 transmission season. Additional peptides also need to be identified and screened for binding to other HLA alleles, to broaden the MHC specificity of the diagnostic reagent or immunopathogenesis tools developed by this approach.
Bioinformatics Analysis
We obtained the NY 1999 WN virus sequence from GenBank (GenBank accession number AF196835) (21). The 3,433 amino acids in the GenBank translation were parsed into 3,424 10-amino acid long frames, each 10 amino acid-long peptide sequence overlapping the previous peptide sequence by nine amino acids. The sequences of these 3,424 decamers were stored in a database.
Each of the peptides in the database was then evaluated by EpiMatrix, a matrix-based algorithm that ranks 9 and 10 amino acid peptides by estimated probability of binding to a selected MHC molecule (22). The estimated binding potential (EBP) is derived by comparing the EpiMatrix score with those of known binders and presumed nonbinders. The EBP describes the proportion of peptides with EpiMatrix scores as high or higher than known binders for a given MHC molecule. Both retrospective and prospective studies of EpiMatrix predictions have confirmed the accuracy of this T cell epitope selection method (22-24). EpiMatrix is available for use by HIV researchers on the TB/HIV Research Laboratory website (http://tbhiv.biomed.brown.edu/) and under collaborative and commercial arrangements with the TB/HIV Research Laboratory and EpiVax, Inc. (Providence, RI), respectively.
Table 1 illustrates the process of selecting candidate B*07 ligands from the WN virus genome. Of six overlapping peptides in the region of the WN virus sequence shown (Table 1), WN virus B7 0019 scored in the same range as known B*07 ligands and HLA B*07-restricted epitopes (EBP 22.49). Therefore, this peptide would be considered the most likely candidate to show binding to HLA B*07 of the six peptides in this illustration.
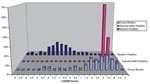
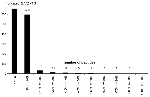
EBPs for the WN virus peptides ranged from >20% (highly likely to bind) to <1% (very unlikely to bind) (Figure 1). We also scored 10,000 random peptides of natural amino acid composition (25) derived from the ExPASy (Expert Protein Analysis System) proteomics server at the Swiss Institute of Bioinformatics (Randseq, http://www.expasy.ch/tools/randseq.html). We compared the HLA B*07 EpiMatrix scores of this set of random peptides with those of a set of >300 known binders (compiled and maintained at EpiVax) and with the scores of the set of WN virus peptides selected for this study (Figure 2).
Selection of Peptides
Peptides with EpiMatrix EBP scores in the range of 7 to 50 are more likely to bind to MHC and stimulate T cells in vitro (23). Peptides with an EBP score >50 are less likely to be immunogenic, although they may bind to B7 in vitro (16,23). Therefore, peptides scoring >50 in the WN virus set were excluded. Based on these criteria, a final set of 22 peptides with EBP scores from 20 to 50 (Table 2) was selected for screening in vitro. We excluded 3,329 WN virus peptides with EBP scores <7, 70 potential B*07 binders with EBP scores >7 and <20, and 3 peptides with scores >50 (Table 2b).
Four of the lowest scoring WN virus peptides (EBP 0.00%, Table 2a) were also selected to test the hypothesis that low scoring peptides derived from WN virus would not bind to HLA B*07 in vitro (predicted nonbinders). One well-defined B*07-restricted epitope, GPGHKARVLA (derived from HIV), was also chosen as a positive control for the assays (26).
Cross-Reactive Analysis
After the EpiMatrix analysis, the Conservatrix tool (EpiVax, Providence, RI) was used to align and compare the WN virus sequences with those of other related flaviviruses (21). In an intermediate step designed to avoid selecting epitopes that may have cross-reactivity with "self," each of the highly selected epitopes was passed through the Blast engine at the National Center for Biotechnology Information, using the BlastiMer tool (EpiVax, Providence, RI). Any sequence that was similar to (i.e., >80% identical to the 10 amino acid WN virus NY99 sequence) a peptide component of equivalent length in the human genome (accessible and published to date) was excluded from the study set.
Peptide Synthesis
Peptides corresponding to the epitope selections were prepared by 9-fluoronylmethoxycarbonyl synthesis on an automated Rainen Symphony/Protein Technologies synthesizer (Synpep, Dublin, CA). The peptides were delivered 90% pure as ascertained by high-performance liquid chromatography, mass spectrophotometry, and UV scan. The peptides were shipped as lyophilized powder, which was diluted in a minimal volume of dimethyl sulfoxide and then diluted to stock concentrations in RPMI 1640 medium (Sigma, St Louis, MO). Peptides that could not be purified to specifications within the study period were not evaluated.
MHC Binding Studies
The T2B7 binding assay method (23,24) relies on the ability of exogenously added peptides to stabilize the class I MHC/beta 2 microglobulin structure on the surface of transporters associated with antigen processing (TAP)-deficient cell lines (27,28). Briefly, the HLA B*07 T2 cell line was prepared for the assay by incubating overnight (16 hours) at 26°C. Before the binding assay, these cells were washed twice in serum-free media. Solutions of the test peptides at three concentrations (final concentrations of 10, 20, and 200 µg/mL in RPMI 1640 (Sigma, St Louis, MO) were plated in triplicate wells of a 96-well, round-bottom assay plate (Becton Dickinson, Lincoln Park, NJ). Sixteen wells containing cells without peptide were included in each plate as background controls.
After 100,000 cells were added to each well, the plates were incubated for 4 hours at 37°C, 5% CO2, followed by centrifugation at 110 x g for 10 minutes at 4°C. The supernatant was discarded, and the remaining cells were resuspended. One hundred L of anti-HLA-B*07 primary antibody-containing hybridoma supernatant was diluted in staining buffer (1:10 dilution of ME1 supernatant produced by HB-119 cell line (ATCC, Rockville, MD) in staining buffer: phosphate-buffered saline [PBS], 5% fetal bovine serum, 0.1% sodium azide) was added to all the wells. Primary antibody was incubated with the peptide-pulsed cells for 30 minutes at 4°C. After washing three times with staining buffer, the cells were resuspended, and 100 µL of a 1:250 dilution of fluorescein isothiocyanate (FITC)-labeled secondary antibody (FITC-labeled Goat F[ab]2 anti-mouse IgG [H+L] [Caltag, Burlingame, CA]) in staining buffer was added to all the wells. The binding studies are predicated on the assumption that the primary antibody recognizes an epitope on the HLA with a configuration that is unchanged by the stabilizing peptide. The plates were incubated for 30 minutes at 4°C, then washed three times with staining buffer. The contents of each well were then resuspended in 200 L of fixing buffer (PBS, 1% paraformaldehyde).
The 16 negative control wells in each plate contained no peptide but did contain cells, primary antibody, and secondary antibody. An additional set of wells was plated with peptide at the highest concentration (200 µg/mL), but no primary antibody was added to the wells as a control for nonspecific secondary antibody binding. One positive control peptide (the known B*07 binder) was tested in triplicate at three concentrations (final concentrations of 10, 20, and 200 µg/mL in RPMI 1640) in each assay plate.
Following fixing, the presence of fluorescent secondary antibody on the surface of T2 cells (gated to the appropriate cell size) was measured at 488 nm on a FACScan flow cytometer (Becton Dickinson). The mean linear fluorescence of 10,000 events was measured and compared with the background fluorescence of cells plated in control wells. The entire assay was repeated 4 times, so that each peptide was tested in a total of 36 wells (triplicate wells, three concentrations, four assays).
The B*07 molecule was considered to be stabilized on the surface of the T2B7 cells if the average of the mean linear fluorescence for the triplicate wells at each concentration of peptide was >10% higher than the average of the 16 negative control wells (and p<0.05 in two-way comparisons by ANOVA). Binding was rated as strong, moderate, weak, or none, based on the number of significantly positive wells by pair-wise ANOVA (Table 3).
The 3,424 decamers derived from the WN virus genome were evaluated by EpiMatrix B*07 and evaluated for match to the stored matrix pattern. Most decamers scored for the entire WN virus genome (by the HLA B*07 scoring matrix) had EBP scores <1% (Figure 1). Figure 2 shows the distribution of HLA B*07 scores of a set of 10,000 random peptides (plotted as their natural logs, to allow better distribution of EBP scores <1), compared with scores for the set of >300 known HLA B*07 binders and with the scores of the selected WN virus peptides. The set of peptides selected for study scored well within the EBP range of the comparison set of >300 known HLA B*07 ligands (Figure 1).
Each peptide in the entire WN virus-NY99 dataset of peptides was scored by EpiMatrix. Ninety-five of the 3,424 decamers had EBP scores >7%. Of these 95 peptides, 20 of the 25 with EBP scores between 20% and 50% (Table 2a) were selected for screening. Three peptides with EBP scores >50 (0001, 0002, 0003) were eliminated from the set of peptides tested because scores in this range are less likely to be B*07 ligands and epitopes (TB/HIV Research Lab and EpiVax, unpub. data). The amino acid sequence of peptide 0012 overlapped substantially with the human genome, and for that reason this peptide was also excluded. Three of the original 25 peptides (0014, 0021, 0025) could not be synthesized to sufficient purity within the study timeframe. Two peptides with EBP scores between 50 and 20 (0016 and 0022) were also not tested because they did not fall within a region of the WN virus genome belonging to a mature WN virus protein, based on information in the GenBank database. Sixteen WN virus peptides remained in the final selection.
The final set of 16 WN virus peptides included two from NS-1, four from NS-2A, five from NS-3, one from NS-4A, five from NS-5, one from env gp E, and one from prM (Table 2a). In addition to these peptides, four predicted nonbinder peptides and a known binder (1291) were also synthesized. Twenty-one peptides were tested in vitro in T2B7 binding assays.
Binding Results
Triplicate wells of peptide at 10, 20, and 200 µg/mL were evaluated in each of the T2 B7 binding assays. Table 3 provides information on the mean fluorescence index for the peptide at 200 µg/mL; the average fold increase over background for the peptide at 10, 20 and 200 µg/mL; and the ANOVA analysis for each pairwise comparison (between fluorescence for cells incubated with one of the concentrations of the study peptide and the fluorescence of the cells in control wells).
Twelve of the 16 study peptides demonstrated consistent binding in the four replicate assays. Of these peptides, four (0017, 0019, 0020, and 0023) stabilized HLA B*07 on the surface of T2B7 cells substantially more often than controls in the four replicate assays (strong binders, Table 3). Two WN virus peptides (0008, 0009) stabilized HLA B*07 to a moderate degree. Six WN virus peptides (0005, 0006, 0013, 0015, 0018, and 0024) were weak binders, and four did not bind.
The positive control peptide, 1291, was tested with each set of peptides. The peptide bound significantly over background (based on ANOVA) in all three assays. Four negative control peptides selected for low EBP scores (3399, 3404, 3411, and 3415, all with scores of 0.0%) did not stabilize T2B7 to a significant degree.
Cross Strain Analysis Results
Peptide 0019, a strong binder, was conserved in all strains of WN virus (100% or 10 of 10 amino acids) and Kunjin virus; it was 80% conserved in JE virus strains (8 of 10 amino acids). Peptides 0017 and 0023, two strong binders, were 100% conserved in all strains of WN virus and Kunjin virus, 80% conserved in JE virus and Murray Valley encephalitis (MVE) virus, and 90% conserved in some strains of dengue. Peptide 0020, the fourth strong binder, was conserved in West Nile and Kunjin (100%), JE virus, MVE virus, and dengue (90%).
The two moderate binders 0008 and 0009 were unique. Peptide 0008 was 100% conserved in West Nile virus strains, different by one amino acid from Kunjin virus (closely related to WN virus), and not conserved in any other flaviviruses. Peptide 0009 was conserved only in WN virus and Kunjin virus (100%) and not conserved in any other related flavivirus strains.
Of the weaker binding peptides (0005, 0006, 0013, 0015, 0018, and 0024), 0005 was 100% conserved across WN, Kunjin, SLE, and Sindbis viruses. Peptide 0006 was 90%-100% conserved in WN, Kunjin, JE, and MVE viruses. Peptide 0013, likewise, was conserved in WN and Kunjin but less well in JE virus (80%). WN virus 0015 was conserved in WN virus, Kunjin (100%), JE virus (90%), MVE virus (80%), SLE virus (90%), and dengue (80%). WN virus 0018 was conserved in WN virus, JE virus (90%), MVE virus (80%), and SLE virus (90%). In contrast, peptide 0024 was 100% conserved in all strains of WN virus (100%, or 10 of 10 amino acids) and Kunjin virus but not conserved in any other virus of the flavivirus group.
Estimated Cost
The 3,329 peptides with EBP scores <7% (3,424 to 95) were considered unlikely to bind to HLA B*07. The EpiMatrix approach reduced the number of candidate peptides by 97% (3,329/3,424). Some researchers have adopted a standard overlapping (OL) approach (constructing a set of 10 amino acid-long peptides overlapping by 5 amino acids covering the entire genome [29]). This strategy (10/5 OL set) would have required the synthesis of 685 decamer peptides for the WN virus genome, more than 7 times the number (95) selected by the EpiMatrix approach.
The cost of synthesizing the 16 putative ligands and four controls (at a cost of $250 per peptide) for this project was $5,000. Synthesizing the entire selected set and four controls would have cost $24,750. Had the standard overlapping peptide approach been used, the cost of synthesizing OL peptides would have been approximately $170,000 ($250 for each of 685 peptides). The cost of synthesizing and mapping the complete overlapping set of peptides representing decamer peptides overlapping by 9 amino acids (3,423 peptides) would be $856,000 (Table 4).
If the WN virus B7 peptides behave as observed in previous studies of HLA B*07 peptide datasets (23; De Groot et al., unpub. data), additional HLA-B7 ligands would be identified (approximately 76%; 72 of the set of 95 WN virus peptides with EBP scores >7). If, by performing more overlapping peptide assays, this larger set of 72 (putative) ligands had been found, the cost per ligand with the OL approach would have been approximately $3,600 per ligand, compared with $617 per ligand for 72 ligands with the EpiMatrix approach. If no epitopes were to be missed, the exhaustive approach could be used at an estimated cost of $18,000 per ligand. This approach would have cost approximately five times more than the OL approach and 30 times more than the EpiMatrix approach.
Time Required for Analysis
Analysis of the WN virus genome and selection of the WN virus peptides was performed during one working week. Selected peptides were obtained in batches over a 4-week period. T2B7 binding assays were performed as the peptides arrived. Overall, the T2 B7 binding assays and data analysis took place over 20 working days, and the entire process from peptide selection to completion of data analysis took 8 weeks. Eliminating delays associated with peptide synthesis would have reduced the time required to 4 weeks.
Using the EpiMatrix approach, we rapidly identified four excellent B*07-restricted T cell epitope candidates for WN virus. Overall, 12 (75%) of 16 selected peptides bound in T2B7 binding assays. These binding results compare favorably with those of other T2B7 binding results for HIV-1 (16,23). Xia Jin et al. tested 29 HIV-1 peptides with EBP scores of 7%, of which 10 (35%) bound to T2B7 cells in vitro and 4 (14%) were subsequently demonstrated to be HLA-B7 restricted CTL epitopes in assays performed with CD8+ T cell lines derived from an HIV-infected patient (23). In a separate study (16) of HLA B*07-restricted peptides, 25 peptides were tested, including a known HLA B*07-restricted epitope (peptide 1291, also used in this study). Nineteen (76%) of 25 peptides were shown to bind to T2B7 cells in vitro, and 60% of the peptides stimulated gamma-interferon release in T cell assays performed with HIV-1-infected patients' cells.
Based on these experiences with EpiMatrix HLA-B7 selection, additional peptides from the original list of 95 WN virus peptides (EBP scores >7) might be expected to bind to HLA B*07 and stimulate T cell responses. If the rest of the WN virus B7 peptides behave as observed in the HIV-1 datasets, 21 to 60 additional HLA-B7 ligands might be identified (76%, or 72 of the set of 95 WN virus peptides with EBP scores >7). This observation is also consistent with estimates of the number of epitopes in a given protein (30). Even at this higher number of total ligands, the cost per ligand of the OL approach would still have been more expensive than the EpiMatrix approach. Furthermore, the exhaustive approach would have cost approximately five times more than the OL (10/5) approach and 30 times more than the EpiMatrix approach. The EpiMatrix approach would also be substantially more rapid than OL or exhaustive testing of overlapping peptides.
EpiMatrix is one of several epitope mapping tools available to researchers, including the tool available at the SYFPEITHI (31) website and the HLA binding prediction tool available on the National Institutes of Health (BIMAS) site (32). Neither of these sites returned exactly the same predictions as EpiMatrix for the WN virus genome; however, no direct comparison was made. Either of these web-based epitope-mapping tools could also accelerate the process of epitope mapping the WN virus genome by the approach described here.
The matrix-based approach used by EpiMatrix developers occasionally results in the selection of peptides that do not fit standard anchor-based and extended anchor motifs such as those available on the SYFPEITHI website. As a result, WN virus peptides selected by the EpiMatrix method and included in this study did not always fit the conventional, anchor-based format of proline in position 2 and leucine or phenylalanine in position 9 (17). For example, the sequence of one weak WN virus binder, AAKKKGASLL, has little in common with published HLA B*07 motifs, illustrating how EpiMatrix is able to prospectively identify ligands that do not necessarily match anchor-based motifs.
Although EpiMatrix appears to provide excellent discrimination between most published HLA B*07 ligands and a set of random peptides (Figure 2), there is still overlap between the lower-scoring published HLA B*07 ligands and the scores of some of the random peptides. Since the universe of HLA B*07 ligands is unknown, some of the set of random peptides could be previously unidentified HLA B*07 ligands. Furthermore, EpiMatrix scored several known HLA B*07 ligands very low, reflecting either inaccuracy of the HLA B*07 matrix or inaccurate reporting of these ligands. Further study of these low-scoring HLA B*07 ligands may improve knowledge of the rules determining HLA B*07 binding.
Epitopes that are specific for WN virus could be used to develop diagnostic tests such as tetramer assays for WN virus (17). The tetramer staining assay relies only on the interaction between the tetramer reagent and T cell receptors on the surface of T cells; it can be performed in <30 minutes on as little as 2 mL of blood. Peptide 0008 was unique to WN virus, with only 8 of 10 amino acids in this sequence conserved in Kunjin virus; the sequence was even less well conserved in other members of the flavivirus family. Peptide 0009 would also be a strong candidate reagent for a diagnostic test, as it was conserved in Kunjin and in many strains of WN virus but not in any other member of the flavivirus family.
The incubation period in humans (i.e., time from infection to onset of disease symptoms) for WN virus encephalitis is usually 5 to 15 days. Antibodies are detectable within 3 to 7 days; however, to confirm infection, antibody assays must be repeated in the acute and convalescent phases. In contrast, recent tetramer-staining studies (33) indicate that cell responses may be detectable 2 to 3 days after acute infection. The initial CTL response to acute infection with a virus, as measured by tetramer technology, can be dramatic. For example, during the acute immune response to lymphocytic choriomeningitis virus (LCMV) in BALB/c mice, 55% of all CD8+ splenocytes are stained with an LCMV-specific tetramer (34). The method is extremely robust and can detect antigen-specific populations at frequencies as low as 1:5,000 CD8+ T cells, or approximately 1:50,000 peripheral blood mononuclear cells (35). Results of the studies performed here suggest that peptides 0008 and 0009, which are relatively specific for WN virus and which score in the range of EpiMatrix scores shown to be compatible with immunogenicity (24), would be reasonable first candidates for the development of a tetramer-based diagnostic reagent for WN virus.
No specific vaccine or antiviral treatment exists for WN virus infection. CTL response will likely be one critical component of the immune response against WN virus. Development of a preventive or therapeutic vaccine against this public health threat would be greatly expedited if the correlates of immune response were determined and appropriate components rapidly incorporated into a vaccine. Epitopes defined by methods such as the one described here are likely to contribute substantially to the development of new research and diagnostic reagents and vaccines for WN virus and other emerging infectious diseases.
Dr. De Groot is director of the TB/HIV Research Laboratory and assistant professor of community health and medicine at Brown University. She is also CEO and President of EpiVax, Inc., a privately owned bioinformatics and vaccine design company in Providence, RI. She trained in informatics and immunology at the National Institutes of Health and is an HIV/AIDS specialist in correctional settings.
References
- Lanciotti RS, Roehrig JT, Deubel V, Smith J, Parker M, Steele K, Origin of the West Nile virus responsible for an outbreak of encephalitis in the northeastern United States. Science. 1999;286:2333–7.PubMedGoogle Scholar
- Wright PJ, Warr HM, Westaway EG. Comparisons by peptide mapping of proteins specified by Kunjin, West Nile and Murray Valley encephalitis viruses. Aust J Exp Biol Med Sci. 1983;61:641–53.PubMedGoogle Scholar
- Anderson JF, Andreadis TG, Vossbrinck CR, Tirrell S, Wakem EM, French RA, Isolation of West Nile virus from mosquitoes, crows, and a Cooper's Hawk in Connecticut. Science. 1999;286:2331–3.PubMedGoogle Scholar
- Rappole JH, Derrickson SR, Hubalek Z. Migratory birds and spread of West Nile virus in the Western Hemisphere. Emerg Infect Dis. 2000;6:319–28.PubMedGoogle Scholar
- Jia XY, Briese T, Jordan I, Rambaut A, Chi HC, Mackenzie JS, Genetic analysis of West Nile New York 1999 encephalitis virus. Lancet. 1999;354:1971–2.PubMedGoogle Scholar
- Aihara H, Takasaki T, Matsutani T, Suzuki R, Kurane I. Establishment and characterization of Japanese encephalitis virus-specific, human CD4(+) T-cell clones: flavivirus cross-reactivity, protein recognition, and cytotoxic activity. J Virol. 1998;72:8032–6.PubMedGoogle Scholar
- Okamoto Y, Kurane I, Leporati AM, Ennis FA. Definition of the region on NS3 which contains multiple epitopes recognized by dengue virus serotype-cross-reactive and flavivirus-cross-reactive HLA-DPw2-restricted CD4+ T cell clones. J Gen Virol. 1998;79:697–704.PubMedGoogle Scholar
- Zeng L, Kurane I, Okamoto Y, Ennis FA, Brinton MA. Identification of amino acids involved in recognition by dengue virus NS3-specific, HLA-DR15-restricted cytotoxic CD4+ T-cell clones. J Virol. 1996;70:3108–17.PubMedGoogle Scholar
- Kurane I, Okamoto Y, Dai LC, Zeng LL, Brinton MA, Ennis FA. Flavivirus-cross-reactive HLA-DR15-restricted epitope on NS3 recognized by human CD4+ CD8- cytotoxic T lymphocyte clones. J Gen Virol. 1995;76:2243–9.PubMedGoogle Scholar
- Shen JT. T-To SS, Schrieber L, King NJ. Early E-selectin, VCAM-1, ICAM-1, and late major histocompatibility complex antigen induction on human endothelial cells by flavivirus and comodulation of adhesion molecule expression by immune cytokines. J Virol. 1997;71:9323–32.PubMedGoogle Scholar
- Johnston LJ, Halliday GM, King NJ. Phenotypic changes in Langerhans cells after infection with arboviruses: a role in the immune response to epidermally acquired viral infection? J Virol. 1996;70:4761–6.PubMedGoogle Scholar
- Johnston LJ, Halliday GM, King NJ. Langerhans cells migrate to local lymph nodes following cutaneous infection with an arbovirus. J Invest Dermatol. 2000;114:560–8.PubMedGoogle Scholar
- Lobigs M, Arthur CE, Mullbacher A, Blanden RV. The flavivirus nonstructural protein NS3 is a dominant source of cytotoxic T cell peptide determinants. Virology. 1994;202:195–201.PubMedGoogle Scholar
- Murali-Krishna K, Ravi V, Manjunath R. Cytotoxic T lymphocytes raised against Japanese encephalitis virus: effector cell phenotype, target specificity and in vitro virus clearance. J Gen Virol. 1994;75:799–807.PubMedGoogle Scholar
- De Groot AS, Bosma A, Chinai N, Frost J, Jesdale BM, Gonzalez MA, From genome to vaccine: in silico predictions, in vivo verification. Vaccine. 2001. In press.PubMedGoogle Scholar
- Altman JD, Moss PA, Goulder PJ, Barouch DH, McHeyzer-Williams MG, Bell JI, Phenotypic analysis of antigen-specific T lymphocytes. Science. 1996;274:94–6.PubMedGoogle Scholar
- Hanke T, Schneider J, Gilbert SC, Hill AV, McMichael A. DNA multi-CTL epitope vaccines for HIV and Plasmodium falciparum: immunogenicity in mice. Vaccine. 1998;16:426–35.PubMedGoogle Scholar
- An LL, Whitton JL. A Multivalent minigene vaccine, containing B-cell, cytotoxic T-lymphocyte, and Th epitopes from several microbes, induces appropriate responses in vivo and confers protection against more than one pathogen. J Virol. 1997;71:2292–302.PubMedGoogle Scholar
- Nardin EH, Calvo-Calle JM, Oliveira GA, Nussenzweig RS, Schneider M, Tiercy JM, A totally synthetic polyoxime malaria vaccine containing plasmodium falciparum B cell and universal T cell epitopes elicits immune responses in volunteers of diverse HLA types. J Immunol. 2001;166:481–9.PubMedGoogle Scholar
- Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402.PubMedGoogle Scholar
- De Groot AS, Jesdale BM, Szu E, Schafer JR, Chicz RM, Deocampo G. An interactive web site providing major histocompatability ligand predictions: application to HIV research. AIDS Res Hum Retroviruses. 1997;13:529–31.PubMedGoogle Scholar
- Jin X, Nixon DF, Roberts CG, Safrit JT, Zhang LQ, Huang YX, Identification of subdominant cytotoxic T lymphocyte epitopes encoded by autologous HIV type 1 sequences, using dendritic cell stimulation and computer-driven algorithm. AIDS Res Hum Retroviruses. 2000;16:67–76.PubMedGoogle Scholar
- Schafer JR, Jesdale BM, George JA, Kouttab NM, De Groot AS. Prediction of well-conserved HIV-1 ligands using a Matrix-based Algorithm, EpiMatrix. Vaccine. 1998;16:1880–4.PubMedGoogle Scholar
- Bairoch A. Amino acid scale: Amino acid composition (%) in the SWISS-PROT Protein Sequence data bank. Release notes for SWISS-PROT release 38 - July 1999. Available at: URL: http://www.expasy.ch/tools/pscale/A.A.SWISS-PROT.html Accessed 01.12.01.
- Goulder PJ, Phillips RE, Colbert RA, McAdam S, Ogg G, Nowak MA, Late escape from an immunodominant cytotoxic T-lymphocyte response associated with progression to AIDS. Nat Med. 1997;3:212–7.PubMedGoogle Scholar
- Ljunggren HG, Stam NJ, Ohlen C, Neefjes JJ, Hoglund P, Heemels MT, Empty MHC class I molecules come out in the cold. Nature. 1990;346:476–80.PubMedGoogle Scholar
- Nijman HW, Houbiers JG, van der Burg SH, Vierboom MP, Kenemans P, Kast WM, Characterization of cytotoxic T lymphocyte epitopes of a self-protein, p53, and a non-self-protein, influenza matrix: relationship between major histocompatibility complex peptide binding affinity and immune responsiveness to peptides. J Immunother. 1993;14:121–6.PubMedGoogle Scholar
- Bond KB, Sriwanthana B, Hodge TW, De Groot AS, Mastro TD, Young NL, An HLA-directed molecular and bioinformatics approach identifies new HLA-A11 HIV-1 subtype E cytotoxic T lymphocyte epitopes in HIV-1 infected Thais. AIDS Res Hum Retroviruses. 2001. In press.PubMedGoogle Scholar
- Woodfolk JA, Sung SS, Benjamin DC, Lee JK, Platts-Mills TA. Distinct human T cell repertoires mediate immediate and delayed-type hypersensitivity to the trichophyton antigen, tri r 2. J Immunol. 2000;165:4379–87.PubMedGoogle Scholar
- Rammensee HG, Bachmann J, Emmerich NP, Bachor OA, Stevanovic S. SYFPEITHI: database for MHC ligands and peptide motifs. [Available at: URL: http://www.uni-tuebingen.de/uni/kxi/]. Immunogenetics. 1999;50:213–9.PubMedGoogle Scholar
- Parker KC, Bednarek MA, Coligan JE. Scheme for ranking potential HLA-A2 binding peptides based on independent binding of individual peptide side-chains. J Immunol. 1994;152:163–75.PubMedGoogle Scholar
- Blattman JN, Sourdive DJ, Murali-Krishna K, Ahmed R, Altman JD. Evolution of the T cell repertoire during primary, memory, and recall responses to viral infection. J Immunol. 2000;165:6081–90.PubMedGoogle Scholar
- Murali-Krishna K, Altman JD, Suresh M, Sourdive DJD, Zajac AJ, Miller JD, Counting antigen-specific CD8 T Cells: A reevaluation of bystander activation during viral infection. Immunity. 1998;8:177–87.PubMedGoogle Scholar
- Callan MF, Tan L, Annels N, Ogg GS, Wilson JD, O'Callaghan CA, Direct visualization of antigen-specific CD8+ T cells during the primary immune response to Epstein-Barr virus in vivo. J Exp Med. 1998;187:1395–402.PubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 7, Number 4—August 2001
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Anne S. De Groot, TB/HIV Research Laboratory, Brown University, Providence, RI 02906, USA; fax: 401-863-1243
Top