Volume 10, Number 4—April 2004
Research
Antimicrobial Resistance Gene Delivery in Animal Feeds
Abstract
Avoparcin, a glycopeptide antimicrobial agent related to vancomycin, has been used extensively as a growth promoter in animal feeds for more than 2 decades, and evidence has shown that such use contributed to the development of vancomycin-resistant enterococci. A cluster that includes three genes, vanH, vanA, and vanX, is required for high-level resistance to glycopeptides. In the vancomycin producer Amycolatopsis orientalis C329.2, homologs of these genes are present, suggesting an origin for the cluster. We found substantial bacterial DNA contamination in animal-feed-grade avoparcin. Furthermore, nucleotide sequences related to the cluster vanHAX are present in this DNA, suggesting that the prolonged use of avoparcin in agriculture led to the uptake of glycopeptide resistance genes by animal commensal bacteria, which were subsequently transferred to humans.
Antimicrobial resistance in bacterial pathogens is a major impediment to successful therapy, and in several instances, bacterial strains have arisen that are refractory to most available antimicrobial treatments (1). Resistance arises by mutation (influencing the target or efflux of the antimicrobial agent) or by the acquisition of resistance genes (encoding antimicrobial or target alteration, or alternate pathways) (2,3). The actual origins of acquired resistance genes are unknown, but environmental microbes, including the strains producing antimicrobial agents, are believed to be important sources (4,5). Substantial genetic and biochemical similarities exist between resistance determinants in antimicrobial agent–producing actinomycetes and resistance genes found in gram-positive and gram-negative pathogens (6–9).
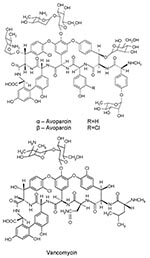
Since vancomycin-resistant enterococci (VRE) were clinically isolated in Europe (1986) and the United States (1987), VRE infections have been reported throughout the world. These infections may be life-threatening because choices for alternative treatment are limited. Concomitant with human use of vancomycin, avoparcin, a closely related glycopeptide antimicrobial agent, has been widely used in Europe and other continents as an animal growth promoter (Figure 1). VRE have been isolated, commonly from pigs and chickens fed avoparcin-containing animal feed, and humans coming into contact with the animals (farm workers, butchers) have been shown to carry VRE (10–12); identical clones have been found (13). The public health concern about the emergence and dissemination of VRE in food animals and the food supply caused the European Union to ban the use of avoparcin in animal feed in 1997. The discontinued use of avoparcin in animal feed has resulted in a reduction in the number of vancomycin-resistant organisms isolated from animals (14,15).
High-level glycopeptide resistance is conferred by a cluster of three genes, vanH, vanA, vanX (the van cluster), plus associated regulatory elements; the cluster is often carried by conjugative transposons (16–18). The vanH gene encodes a D‑lactate dehydrogenase that provides the requisite D‑lactate. vanX encodes a highly specific DD-peptidase that cleaves only D-Ala-D-Ala produced endogenously while leaving D-Ala-D-Lac intact. The third gene, vanA, encodes an ATP-dependent D-Ala-D-Lac ligase. Replacement of D-Ala-D-Ala by D-Ala-D-Lac in the bacterial cell wall results in a thousandfold reduction in the binding of glycopeptide antimicrobial agents to their peptidoglycan target (19). Studies have demonstrated the presence of vanHAX homologs, such as vanH-ddlN-vanX (Figure 2), in actinomycete strains producing glycopeptides, and strong structural and functional similarity exists between the various homologs and the van cluster of VRE (8,9). Some researchers have proposed that the vanH, vanA, and vanX genes of hospital enterococci may have been acquired en bloc from the actinomycetes (8). Related vanHAX gene clusters have been identified in Paenibacillus spp. by Patel and coworkers, indicating another possible source of the van cluster (20). Regardless of the microbial source, the feeding of crude antimicrobial preparations to animals is plausible as a delivery process for transferring the cognate antimicrobial resistance genes between producing strains and the commensal bacteria of animals (21); the concomitant selection for resistance would ensure the survival of rare resistant strains. We provide evidence that a DNA-encoding homolog of the van cluster is a contaminant of feed-grade avoparcin and propose that animal use both created and selected for glycopeptide-resistant strains. The emergence of vancomycin-resistant Staphylococcus aureus (VRSA) is a recent sequela to this train of events involving the van gene clusters (22).
DNA Extraction from Avoparcin
A suspension (0.7 mL) of avoparcin (Roche, Sydney, Australia) in H2O (100 μg/mL) was centrifuged in a 1.5-mL Eppendorf tube for 6 min and the supernatant, after being shaken with 1 volume of phenol:chloroform:isoamyl alcohol (25:24:1), was centrifuged at 16,000 x g for 3 min. The aqueous phase was subjected to two additional phenol-chloroform extractions. The nucleic acid in the pooled aqueous fractions was precipitated with ethanol; the pellet was recovered by centrifugation and further purified by using a GeneClean spin kit (BIO101) and resuspended in 100 μL of double-distilled H2O. The DNA concentration was measured with a fluorometer (Model TKO100, Hoefer Scientific Instruments, San Francisco, CA).
PCR Amplification of 16S rDNA Sequences
Primers 16S 440F and 16S 1491R (Table) were designed to amplify partial 16S rDNA sequences. The polymerase chain reaction ((PCR) contained 2 mM MgCl2, 0.16 mM dNTP, 0.4 μL of each primer, Taq polymerase (1 U), 3‑15 ng template, and 5% dimethyl sulfoxide (DMSO). PCR was done in a MiniCycler (MJ Research, Waltham, MA) by using the following program: 96°C, 3 min; 96°C, 30 s; 60°C, 45 s; 72°C, 1 min 30 s; 35 cycles; and 72°C, 10 min.
PCR Amplification of vanH, ddlN, and vanX Sequences
Different combinations of PCR primers (9) were used to amplify the entire van cluster (Table). Reaction conditions were as described previously.
Cloning of vanH, ddlN, vanX, and Partial 16S rDNA Genes
PCR products were cloned by using vector pCR 2.1-TOPO (Invitrogen, Burling, Ontario, Canada) according to the manufacturer’s instructions, and the insertion size was confirmed by a second PCR. Plasmid DNA was extracted by using the Concert rapid plasmid miniprep system (Invitrogen).
DNA Sequence Analysis
Cycle sequence reactions were carried out with a BigDye terminator DNA sequencing kit (Applied Biosystems, Foster City, CA) with plasmid DNA templates. The cycle sequence program was as follows: 96°C, 1 min; 96°C, 30 s; 50–60°C (dependent on different primers and fragments), 15 s; 60°C, 4 min, for 25 cycles. Excess oligos and dyes were removed by using CentriSep spin columns (Princeton Separations, Aldelphia, NJ). Reaction products were sequenced by the Nucleic Acid and Protein Service, University of British Columbia, using an ABI PRISM 377 sequencer. Sequences were analyzed by using the standard nucleotide-nucleotide BLAST program (National Center for Biotechnology Information, Bethesda, MD). and comparisons were carried out by using CLUSTAL W (European Bioinformatics Institute, Cambridge, UK).
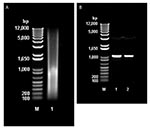
Direct extraction of avoparcin powder with phenol/chloroform/isoamyl alcohol provided substantial amounts of DNA (30.5 μg/g of avoparcin) (Figure 3A). PCR amplification of the DNA with oligonucleotide primers specific for a region of streptomycete 16S rRNA gave a single amplicon (Figure 3B), which was sequenced and shown to be 16S closely related to that of Amycolatopsis coloradensis, the producer of avoparcin. Figure 3B shows similarities between the 16S rRNA of species that produce glycopeptide antimicrobial agents.
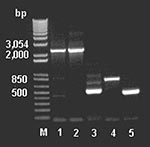
To examine for the presence of genes involved in glycopeptide resistance from the antimicrobial agent–derived DNA, we used the DNA primers described by Marshall et al. (9). The amplicons (Figure 4) were cloned, sequenced, and assembled, indicating a van-like cluster closely related to that found in A. orientalis and Streptomyces toyocaensis. Control reactions run without added template were negative.
The genes encoded three putative proteins showing >50% amino acid identity to the Van H, A, and X proteins of VRE (Figure 2). All of the clusters have translational overlaps between the vanA and vanX genes and their homologs, suggesting cotranslational regulation of expression. This finding clearly implies that the van cluster must be transferred and acquired in toto from any source organism.
We suggest that the use of crude avoparcin preparations in animal feeds from 1975 to 1996 was the origin of the vanHAX cluster in the genesis of VRE (and possibly that found in VRSA) (22,23). Large amounts of avoparcin were used in animal feed; in Denmark, for example, total vancomycin use in 1994 amounted to 24 kg, whereas avoparcin use in animals was 24,000 kg (24). During their entire lives, broiler chickens received 15 mg/kg and pigs 20–40 mg/kg of antimicrobial agentin their feed. Each pig was fed 5–10 g of the crude drug for its life span and, consequently, received a steady dose of DNA encoding vancomycin resistance. In Europe, an estimated 100 mg of antimicrobial agents are used in animal feed for the production of 1 kg of meat for human consumption. We believe that this regimen would have favored the selection and maintenance of rare bacterial transformants carrying the resistance genes. If one bears in mind that large numbers of pigs and chickens were exposed to the antimicrobial agent, the probability of gene pick-up by bacterial commensals in the animal gastrointestinal tract would be favored, and once incorporated into a gut commensal genome, further dissemination would have followed under antimicrobial selection. The finding that organization of the van cluster in contaminating DNA of the feed is identical to that in VRE, with overlapping reading frames typical of translational coupling of gene expression between the vanA and vanX homologs (9), reinforces this supposition.
The mechanism by which a van cluster becomes functionally integrated into bacteria is not known. We propose that intestinal bacteria were the original recipients of the DNA; many of the resident strains are known to be competent for DNA uptake (25,26). However, mere uptake is not sufficient for function, and the actinomycete genes differ from VRE genes in their G+C content (approximately 65% vs. 50%) and codon usage. Given the enormous complexity of bacterial populations in the mammalian gastrointestinal tract (27,28), we assume that a variety of intestinal species may have incorporated the resistance-encoding DNA; expression (at low levels) would have been rare, depending on the compatibility of the van genes with the transcription and translation system of the host. Under constant antimicrobial selection pressure, translationally competent sequences would have developed by mutation; this would not necessarily have occurred in enterococci. Nonetheless, the conversion (evolution) of the actinomycete genes into functional enterococcal genes likely would have required many generations of growth under constant selection, and any intermediate stages in this process are a matter of speculation; however, once established on conjugative transposons, the genes would be readily disseminated (17,29). A number of similar van clusters have been identified in different bacterial species, and whether these evolved independently or by divergent evolution is unknown.
The finding of resistance genes in crude antimicrobial products intended to be fed to animals adds to the already strongly voiced opinion that use of antimicrobial agents in this way constitutes a serious public health concern and further emphasizes the need for prohibiting the use in animal feed of all antimicrobial agents that are employed in human therapy. This ban should include structurally or biologically related antimicrobial agents and the use of any compound with the potential to select for cross-resistance to another antimicrobial agent (15,30). The use of avoparcin in Denmark was prohibited in 1995 and in the European Union in 1997. Subsequently, several other antimicrobial growth promoters were banned (31,32). However, the United States and Canada permit the use of many such products, including penicillin, tetracycline, macrolides, and sulfonamides. Nonhuman applications of antimicrobial agents, such as in agriculture and aquaculture, should employ only chemically and biologically distinct classes of compounds developed specifically for that purpose. Clearly such measures should be combined with a requirement for rational and prudent measures for antimicrobial use in the human population.
Many antimicrobial agents (or their close structural relatives) have been used extensively as animal-feed additives. In almost all cases, crude antimicrobial preparations are used, and thus the antimicrobial agent acts as a carrier for its cognate resistance genes. These delivery systems provide the opportunity for resistant strains of bacteria to evolve and so create an enormous gene pool for antimicrobial resistance determinants in the environment.
Ms. Lu is the senior technician and administrator of the Davies Laboratory, Department of Microbiology and Immunology, University of British Columbia. Her principal interests are the characterization of antimicrobial agent–resistant bacteria and their mechanisms of resistance. She is currently working on a reporter system to classify antimicrobial activity.
Acknowledgments
We thank Peter Collignan for his interest and assistance and Dorothy Davies for manuscript preparation.
This work was supported by the Canadian Bacterial Diseases Network.
References
- Levy SB. The antibiotic paradox: how miracle drugs are destroying the miracle. New York: Plenum Press, 1992.
- Davies J. Inactivation of antibiotics and the dissemination of resistance genes. Science. 1994;264:375–82. DOIPubMedGoogle Scholar
- Chadwick DJ, Goode J, eds. Antibiotic resistance: origins, evolution, selection and spread. (Ciba Foundation Symposium 207). Chichester (UK): Wiley; 1997.
- Benveniste R, Davies J. Aminoglycoside antibiotic-inactivating enzymes in actinomycetes similar to those present in clinical isolates of antibiotic-resistant bacteria. Proc Natl Acad Sci U S A. 1973;70:2276–80. DOIPubMedGoogle Scholar
- Piepersberg W, Distler J, Heinzel P, Perez-Gonzalez J-A. Antibiotic resistance by modification: many resistance genes could be derived from cellular control genes in actinomycetes. A hypothesis. Actinomycetologica. 1988;2:83–98. DOIGoogle Scholar
- Shaw KJ, Rather PN, Hare RS, Miller GH. Molecular genetics of aminoglycoside resistance genes and familial relationships of the aminoglycoside-modifying enzymes. Microbiol Rev. 1993;57:138–63.PubMedGoogle Scholar
- Kirby R. Evolutionary origin of aminoglycoside phosphotransferase resistance genes. J Mol Evol. 1990;30:489–92. DOIPubMedGoogle Scholar
- Marshall CG, Broadhead G, Leskiw BK, Wright GD. D-Ala-D-Ala ligases from glycopeptide antibiotic-producing organisms are highly homologous to the enterococcal vancomycin-resistance ligases VanA and VanB. Proc Natl Acad Sci U S A. 1997;94:6480–3. DOIPubMedGoogle Scholar
- Marshall CG, Lessard IAD, Park I-S, Wright GD. Glycopeptide antibiotic resistance genes in glycopeptide-producing organisms. Antimicrob Agents Chemother. 1998;42:2215–20.PubMedGoogle Scholar
- Stobberingh E, van den Bogaard A, London N, Driessen C, Top J, Willems R. Enterococci with glycopeptide resistance in turkeys, turkey farmers, turkey slaughterers, and (sub)urban residents in the south of The Netherlands: evidence for transmission of vancomycin resistance from animals to humans? Antimicrob Agents Chemother. 1999;43:2215–21.PubMedGoogle Scholar
- Witte W. Medical consequences of antibiotic use in agriculture. Science. 1998;279:996–7. DOIPubMedGoogle Scholar
- Hammerum AM, Fussing V, Aarestrup FM, Wegener HC. Characterization of vancomycin-resistant and vancomycin-susceptible Enterococcus faecium isolates from humans, chickens and pigs by RiboPrinting and pulsed-field gel electrophoresis. J Antimicrob Chemother. 2000;45:677–80. DOIPubMedGoogle Scholar
- Manson JM, Keis S, Smith JM, Cook GM. A clonal lineage of VanA-type Enterococcus faecalis predominates in vancomycin-resistant enterococci isolated in New Zealand. Antimicrob Agents Chemother. 2003;47:204–10. DOIPubMedGoogle Scholar
- Aarestrup FM, Seyfarth AM, Emborg H-D, Pedersen K, Hendriksen RS, Bager F. Effect of abolishment of the use of antimicrobial agents for growth promotion on occurrence of antimicrobial resistance in fecal enterococci from food animals in Denmark. Antimicrob Agents Chemother. 2001;45:2054–9. DOIPubMedGoogle Scholar
- Bonten MJM, Willems R, Weinstein RA. Vancomycin-resistant enterococci: why are they here, and where do they come from? Lancet Infect Dis. 2001;1:314–25. DOIPubMedGoogle Scholar
- Arthur M, Courvalin P. Genetics and mechanisms of glycopeptide resistance in enterococci. Antimicrob Agents Chemother. 1993;37:1563–71.PubMedGoogle Scholar
- Pootoolal J, Neu J, Wright GD. Glycopeptide antibiotic resistance. Annu Rev Pharmacol Toxicol. 2002;42:381–408. DOIPubMedGoogle Scholar
- Tomita H, Pierson C, Lim SK, Clewell DB, Ike Y. Possible connection between a widely disseminated conjugative gentamicin resistance (pMG1-like) plasmid and the emergence of vancomycin resistance in Enterococcus faecium. J Clin Microbiol. 2002;40:3326–33. DOIPubMedGoogle Scholar
- Walsh CT, Fisher SL, Park I-S, Prahalad M, Wu Z. Bacterial resistance to vancomycin: five genes and one missing hydrogen bond tell the story. Chem Biol. 1996;3:21–8. DOIPubMedGoogle Scholar
- Patel R. Enterococcal-type glycopeptide resistance genes in non-enterococcal organisms. FEMS Microbiol Lett. 2000;185:1–7. DOIPubMedGoogle Scholar
- Webb V, Davies J. Antibiotic preparations contain DNA: a source of drug resistance genes? Antimicrob Agents Chemother. 1993;37:2379–84.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. Staphylococcus aureus resistant to vancomycin-United States, 2002. JAMA. 2002;288:824–5. DOIPubMedGoogle Scholar
- Chang S, Sievert DM, Hageman JC, Boulton ML, Tenover FC, Downes FP, Infection with vancomycin-resistant Staphylococcus aureus containing the vanA resistance gene. N Engl J Med. 2003;348:1342–7. DOIPubMedGoogle Scholar
- Wegener HC. Historical yearly usage of glycopeptides for animals and humans: The American-European paradox revisited. Antimicrob Agents Chemother. 1998;42:3049.PubMedGoogle Scholar
- Bertolla F, Simonet P. Horizontal gene transfers in the environment: natural transformation as a putative process for gene transfers between transgenic plants and microorganisms. Res Microbiol. 1999;150:375–84. DOIPubMedGoogle Scholar
- Lorenz MG, Wackernagel W. Bacterial gene transfer by natural genetic transformation in the environment. Microbiol Rev. 1994;58:563–602.PubMedGoogle Scholar
- Shoemaker NB, Vlamakis H, Hayes K, Salyers AA. Evidence for extensive resistance gene transfer among Bacteroides spp. and among Bacteroides and other genera in the human colon. Appl Environ Microbiol. 2001;67:561–8. DOIPubMedGoogle Scholar
- Hooper LV, Gordon JI. Commensal host-bacterial relationships in the gut. Science. 2001;292:1115–8. DOIPubMedGoogle Scholar
- Grohmann E, Muth GW, Espinosa M. Conjugative plasmid transfer in gram-positive bacteria. Microbiol Mol Biol Rev. 2003;67:277–301. DOIPubMedGoogle Scholar
- Butaye P, Devriese LA, Haesebrouck F. Antimicrobial growth promoters used in animal feed: effects of less well known antibiotics on gram-positive bacteria. Clin Microbiol Rev. 2003;16:175–88. DOIPubMedGoogle Scholar
- World Health Organization. WHO global principles for the containment of antimicrobial resistance in animals intended for food. Report of a WHO consultation 5–9 June 2000, Geneva Switzerland. WHO/CDS/CSR/APH/2000.4. Geneva: The Organization; 2000.
- European Commision, Scientific Steering Committee. Opinion of the Scientific Steering Committee on antimicrobial resistance. Brussels: European Commission, Directorate-General DGXXIV; 1999.
Figures
Table
Cite This ArticleTable of Contents – Volume 10, Number 4—April 2004
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Julian Davies, Department of Microbiology and Immunology, #300 – 6174 University Boulevard, Vancouver, B.C., Canada V6T 1Z3; fax: 604-822-6041
Top