Volume 11, Number 9—September 2005
Research
Fluoroquinolone-resistant Escherichia coli, Indonesia
Abstract
In a recent, population-based survey of 3,996 persons in Indonesia, fluoroquinolone (FQ)-resistant Escherichia coli was prevalent in the fecal flora of 6% of patients at hospital admission and 23% of patients at discharge, but not among healthy relatives or patients visiting primary healthcare centers (2%). Molecular typing showed extensive genetic diversity with only limited clonality among isolates. This finding suggests that independent selection of resistant mutants occurs frequently. FQ-resistant isolates exhibited a higher rate of spontaneous mutation, but sparser virulence profiles, than FQ-susceptible isolates from the same population. The resistant isolates belonged predominantly to phylogenetic groups A (57%) and B1 (22%) but also to the moderately virulent group D (20%). Hypervirulent strains from the B2 cluster were underrepresented (1%). Because FQ-resistant E. coli can cause disease, especially nosocomial infections in immunocompromised patients, spread of such strains must be stopped.
Escherichia coli is a common constituent of the gastrointestinal flora of most vertebrates, including humans, and may be isolated from a variety of environmental sources. While most strains are nonpathogenic, certain ones can cause a variety of intestinal and extraintestinal infections. Pathogenicity is largely determined by gene-encoding virulence factors (VFs), such as adhesins, toxins, and polysaccharide surface coatings (1). Phylogenetic analysis showed that most E. coli strains fall into 4 main phylogenetic groups, designated A, B1, B2, and D (2). E. coli strains that cause extraintestinal infections derive predominantly from group B2 and, to a lesser extent, group D. Strains of groups A and B1 represent most commensal strains and are largely devoid of virulence determinants (3). Although strains harboring a robust extraintestinal VF repertoire cluster predominantly in groups B2 and D, isolates within each phylogenetic group can be further classified as extraintestinal pathogenic E. coli (ExPEC) or non-ExPEC depending on whether specific virulence traits are present (4,5).
The fluoroquinolones (FQs) are potent antimicrobial agents used for the treatment and prophylaxis of infections caused by gram-negative bacteria, including E. coli. FQ-resistant E. coli has been reported increasingly during the last decade in both the hospital environment and the community, which may ultimately limit the utility of these broad-spectrum agents (6–8). Moreover, FQ-resistant E. coli strains often show resistance to other drugs, such as ampicillin, tetracycline, chloramphenicol, trimethoprim-sulfamethoxazole, and gentamicin (7,9). Recent reports have suggested that clinical FQ-resistant E. coli actually tends to be less virulent than susceptible isolates. FQ-resistant E. coli from hospitalized Dutch patients derived predominantly from the low-virulence phylogenetic groups A and B1. None of the 13 invasive isolates derived from phylogenetic group B2 (10). In addition, evidence suggests that clinical FQ-resistant E. coli isolates from humans in Iowa were associated with a shift toward non-B2 phylogenetic groups and to a lower overall virulence genotype (4). FQ resistance may also be associated with strains that intrinsically have a higher overall mutation rate, since the resistance to FQs in E. coli involves the accumulation of multiple spontaneously occurring point mutations in several genes (9,11). These associations, however, may depend on the strains' geographic or clinical origin.
In our study, we investigated these putative associations in a well-defined collection of isolates from Indonesia. A population-based survey of ≈4,000 people in 2 cities on the island of Java (Surabaya and Semarang) was initiated in 2000 by the Antimicrobial Resistance in Indonesia, Prevalence and Prevention study group to investigate the level of carriage of resistant microorganisms. FQ-resistant E. coli was prevalent in the fecal flora of 6% of patients at hospital admission and 23% of patients at discharge but not among healthy relatives or patients visiting primary healthcare centers (2% in both groups) (ES Lestari, unpub. data). In our study, we analyzed these FQ-resistant E. coli isolates to elucidate their molecular epidemiology and virulence. To define clonal relatedness, we performed enterobacterial repetitive intergenic consensus (ERIC) polymerase chain reactions (PCR). The phylogenetic background and virulence profile of these isolates were determined by PCR methods and compared with similar data for FQ-susceptible E. coli isolated from the same population. Finally, we examined the link between FQ resistance and the intrinsic mutation rate.
Strains
The study group program surveillance was initiated to determine the prevalence of antimicrobial resistance in Indonesia. Four different groups of persons in Surabaya and Semarang were studied for carriage of resistant microorganisms in their stools. The 4 groups were patients on the day of admission to the hospital (group 1), patients on the day of discharge after >5 days of hospitalization (group 2), patients visiting a primary healthcare center (group 3), and healthy relatives or household members of group 1 patients (group 4). In groups 1 and 2, rectal swabs were taken from patients in the internal medicine, surgery, gynecology/obstetrics, or pediatrics departments. The specimens were collected from July to October 2001 in Surabaya and from January to May 2002 in Semarang. Further details on the methods of culturing will be published elsewhere. A total of 5,535 E. coli isolates from 3,284 patients were cultured. Antimicrobial susceptibility testing was performed for 1 isolate per patient. The overall by-isolate prevalence of resistance to ciprofloxacin as determined by disk diffusion was 8%. The prevalence of resistance was highest among patients on the day of discharge (18% in Surabaya and 27% in Semarang) and lowest among patients visiting primary healthcare centers and among family members of patients admitted to the hospital (2% in both groups). The prevalence of FQ-resistant E. coli among patients who were tested on the day of admission was 8% in Surabaya and 4% in Semarang. We studied 196 FQ-resistant isolates in more detail. Seventy-five (38%) of these were from Surabaya (19, 48, 4, and 4 isolates from stated population groups 1, 2, 3, and 4, respectively) and 121 (62%) from Semarang (13, 92, 11, and 5, respectively). The FQ-resistant isolates were recovered from patients from all 4 hospital departments in both cities. In Semarang, 43% of these isolates were from surgery departments and 41% were from internal medicine departments. In Surabaya, 43% of the isolates were from the internal medicine department. All 196 ciprofloxacin-resistant E. coli and 200 ciprofloxacin-susceptible E. coli (20 randomly chosen isolates from groups 1, 2, and 3 and 40 from group 4, from each city) were confirmed by Vitek 2 (bioMérieux, Marcy-l'Etoile, France) according to the manufacturer's instructions and included in the molecular analyses.
DNA Isolation
Bacterial DNA was isolated by using the MagNA Pure LC with the MagNA Pure LC DNA Isolation Kit III for bacteria and fungi (standard protocol; Roche Molecular Biochemicals, Mannheim, Germany). DNA concentration was assessed spectophotometrically. Samples were frozen at –20°C until used.
Bacterial Typing by ERIC-PCR
ERIC-PCR was conducted with primers ERIC-1R and ERIC-2 as described previously (8,12,13). The amplification products were subjected to electrophoresis in a 1% agarose gel and were stained with ethidium bromide (50 μg/mL). The ERIC-PCRs were performed by 1 technician within 1 month. Profiles were visually analyzed by 2 microbiologists. Single-band differences in profiles among strains led to the definition of separate genotypes. Ambiguous isolates were retested and analyzed by 2 other microbiologists.
Phylogenetic Analysis and Virulence Typing
Isolates were assigned to 1 of 4 main E. coli phylogenetic groups (A, B1, B2, and D) according to an established triplex PCR assay, in which the 4 phylogenetic groups yield distinct combinations of 3 possible PCR products, chuA (heme transport), yjaA (unknown function gene from E. coli K-12 genome), and TSPE4.C2 (anonymous fragment identified by subtractive hybridization) (2,14). All isolates were screened for 5 ExPEC-defining virulence markers, papA/papC, sfa/focDE, afa/draBC, kpsM II, and iutA. Based on previous statistical analyses of similar data, from collections within which each isolate's ExPEC status could be inferred based on ecologic source or experimental virulence, isolates were classified as ExPEC if positive for >2 of these 5 defining virulence markers (4). All isolates were also screened for hlyD (hemolysin), another ExPEC-associated VF. Subsequently, all isolates that satisfied molecular criteria for ExPEC were screened for 32 additional virulence markers3. These virulence genes were detected by a combination of multiplex PCR and dot-blot hybridization with primers specific for internal or flanking sequences and probes generated and labeled with these primers; this method was previously validated by using dot-blot hybridization with defined control strains (15). A VF score was calculated for each strain as the sum of all VF genes for which the strain tested positive. In all of these PCR assays, the identity of the PCR products was deduced by comparing their size to molecular size standards in ethidium bromide–stained agarose gels. Appropriate positive and negative controls were included in each run.
Mutation Rate Analysis
The mutation rate was determined for 20 randomly selected isolates from phylogenetic group A (10 FQ-susceptible and 10 FQ-resistant) by monitoring the isolates' capacity to generate mutations conferring resistance to rifampin, as described previously (9,16). Forty independent cultures of each of the 20 strains were set up in Luria broth. After overnight incubation, equal concentrations of cultures were suspended in 0.85% NaCl. The suspensions were spread on Luria agar plates containing 100 μg/mL rifampin and incubated overnight. For each strain, the proportion of cultures giving no resistant mutants was used to calculate the mutation rate per cell per generation according to the fluctuation test of Luria and Delbrück. To avoid confounding by variation in phylogenetic background, only phylogenetic group A isolates were investigated. For comparisons of results, we used the relative mutation rate, which was defined as the rate relative to the rate for E. coli strain Nu14 (5 × 10–9 per cell per generation) (9).
Statistical Analysis
All data were analyzed by using the statistic software packages SPSS version 10.0 (SPSS, Chicago, IL, USA) and EpiInfo version 5.00 (Centers for Disease Control and Prevention, Atlanta, GA, USA). Chi-square or Fisher exact tests (2-tailed) were used when appropriate for comparisons of proportions. Comparisons involving VF scores and relative mutation rates were analyzed by using the Mann-Whitney U test. The criterion for statistical significance was a p value <0.05.
Spread of FQ-resistant E. coli
Genetic heterogeneity among the 196 FQ-resistant E. coli was assessed by ERIC-PCR. We documented 158 different patterns, designated types 1–158, which indicated a genetically diverse collection of strains. Twenty pairs of isolates with identical profiles were identified, and 9 distinct multiple-isolate clones were represented by isolates from 3 patients each. The limited number of shared genotypes was mainly recovered from group 2 patients, i.e., patients at the time of discharge from the hospital, 49 (73%) of 67 isolates. Among the total number of 140 isolates from group 2, we identified 119 different ERIC-PCR profiles.
Type 37 occurred in 3 patients from the internal medicine department in Surabaya; all 3 patients were present within this department on the same day. The finding of this unique isolated cluster can be explained by patient-to-patient transmission or a nonpatient-associated environmental source. This explanation was not further examined in this study. Type 90 was isolated from 2 patients on the day they were discharged from the internal medicine department in Semarang. Samples were collected on consecutive days. An isolate with an identical ERIC-PCR pattern was found in the same period in the same hospital in a pediatric patient at discharge. No further obvious clustering in time and place was observed among isolates from the 9 multiple-strain clusters.
Phylogenetic Analysis
PCR-based phylotyping showed that the 200 FQ-susceptible isolates were predominantly from phylogenetic groups A (52%) and B1 (30%) (Table 1). The 196 FQ-resistant isolates also mainly derived from phylogenetic groups A (57%) and B1 (22%), but some derived from the moderately virulent phylogenetic group D (20%). Hypervirulent strains from the B2 cluster were underrepresented (1%). Eighteen (67%) of the 27 isolates from the 9 distinct clones that were represented by 3 isolates each belonged to group A.
Table 1 shows that the resistant isolates were significantly depleted for phylogenetic group B2 and enriched for group D, when compared with the susceptible isolates. These shifts in phylogenetic distribution were significant both overall and specifically in Semarang, whereas a similar but nonsignificant trend was observed in Surabaya.
The phylogenetic distribution of all 396 isolates among the 2 cities was highly similar (data not shown). Comparisons of the distributions among the 4 population groups showed that group D isolates were more often obtained from patients sampled on the day of discharge than from other population groups (37 [21%] of the 180 group 2 isolates belonged to group D versus 25 [12%] of the 216 nongroup 2 isolates, p = 0.01). Stratification showed, however, that this association was due to the excess prevalence of FQ-resistant group D isolates among the group 2 patients. Furthermore, B2 isolates were significantly more prevalent in group 3, patients visiting public healthcare centers (7 [13%] of the 55 group 3 isolates belonged to group B2 versus 10 [3%] of the 341 nongroup 3 isolates, p = 0.004).
Virulence Typing
All E. coli isolates were tested for a set of virulence factors to allow an inference as to their pathogenic potential. The overall prevalence of the 5 defining ExPEC VFs ranged from 2% (sfa/focDE) to 33% (iutA) (Table 1). The FQ-resistant isolates were significantly depleted for papA, papC, sfa/focDE, afa/draBC, hlyD, and kpsM II (Table 1), when compared with the susceptible isolates. Accordingly, 40 (20%) FQ-susceptible E. coli isolates, but only 4 (2%) FQ-resistant isolates (2%), were classified as ExPEC, as they exhibited >2 of the 5 key ExPEC VFs (p<0.001). Thus, FQ resistance was associated with reduced inferred virulence. All FQ-resistant E. coli isolates from the 9 distinct clones that were represented by 3 patients each were found to be non-ExPEC.
The distribution of the 6 screening VFs was also analyzed in relation to the 4 phylogenetic groups (Table 2). Each VF was broadly distributed, occurring in >3 phylogenetic groups. However, papA, papC, kpsM II, hlyD, and sfa/focDE were all significantly associated with phylogenetic group B2. Accordingly, 53% of the phylogenetic group B2 isolates qualified as ExPEC versus 9% of the non-B2 isolates (p<0.001) (Tables 2 and 3).
The 44 ExPEC isolates were studied in more detail (Table 3). The ExPEC isolates derived mainly from phylogenetic groups A (36%) and B1 (32%), with the 4 FQ-resistant ExPEC isolates belonging to groups A (n = 2) and D (n = 2). Many (36%) ExPEC isolates originated from patients on the day of discharge. Both of the FQ-resistant ExPEC isolates from group 2 were from patients in the surgical ward in Semarang. Again, no evidence for clonality was seen. The 4 resistant ExPEC isolates exhibited sparse VF profiles when compared with the susceptible ExPEC isolates. These isolates lacked classic ExPEC VFs such as focG, hlyD, and cnf1. Four other VFs, iha, sat, fyuA, and malX, were more prevalent among susceptible, rather than resistant, ExPEC isolates. Only ibeA was more prevalent among the resistant isolates. The VF iutA was detected in all FQ-resistant ExPEC isolates and in 27 (68%) of the 40 FQ-sensitive isolates. This difference was not significant. Aggregate VF scores were lower among FQ-resistant ExPEC isolates (median 6, range 4–8) than among the 40 FQ-susceptible ExPEC isolates (median 10, range 3–16, p = 0.024).
Mutation Rate
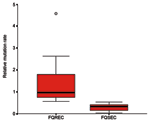
The link between mutation rate and resistance to FQs was studied, as the rate of mutation accumulation might be a factor in the development of FQ resistance. The 10 FQ-susceptible isolates had relative mutation rates of <0.52 (median rate 0.32, range 0.03–0.52), whereas the 10 FQ-resistant E. coli exhibited relative mutation rates of >0.55 (median rate 0.97, range 0.55–4.58) (p<0.001) (Figure).
In this study, we investigated the epidemiology and virulence characteristics of FQ-resistant E. coli collected during a large, population-based survey of ≈4,000 people in 2 cities in Indonesia (Surabaya and Semarang). The overall prevalence of resistance to ciprofloxacin was 8%, but in the fecal flora of patients at time of discharge from the hospital the prevalence was 23%.
Dissemination of FQ-resistant E. coli and Mutation Rate
Three possible explanations for the high prevalence of FQ-resistant E. coli among patients that had been hospitalized for >5 days must be considered: transferable resistance, clonal spread, and mutation-based selection of resistance fostered by the use of antimicrobial agents. Transferable plasmid-mediated quinolone resistance has been described recently in E. coli from China (17). Wang et al. found that 6 (8%) of 78 ciprofloxacin-resistant E. coli strains from a hospital in Shanghai contained qnr. However, from the present study we cannot draw any conclusion about the contribution of this mechanism in Indonesia. As for clonal spread, molecular typing showed extensive genetic diversity among FQ-resistant isolates in Indonesia. We identified a few distinct multiple-isolate clones in the hospital environment. Although all these clonal strains were shown to be non-ExPEC, they may still pose a health threat, especially to immunocompromised patients in hospital settings. Nosocomial outbreaks of infections caused by disseminating FQ-resistant clones have already been described (8). However, in our study, limited clonality among isolates was found, which suggests that other factors contribute more to the high prevalence of FQ-resistant E. coli among hospitalized patients.
To determine whether mutation-based resistance fostered by selection pressure contributed to the prevalence of FQ-resistant E. coli in Indonesia, we performed a mutation rate analysis of selected isolates. We found a strong correlation between resistance and an elevated mutation rate. This finding agrees with a recent report from Komp Lindgren et al., in which high mutation rates of E. coli strains from urinary tract infections were strongly associated with FQ resistance (9). To demonstrate that this mutation-based resistance was selected for by the use of FQs, we must know the consumption figures of the quinolones. In other reports, evidence suggests that the use (and misuse) of ciprofloxacin in human and animal medicine may predispose to an increase in infections with resistant E. coli (8). As information on the use of FQs in Indonesia is currently not available, we cannot draw any conclusions on a potential link between antimicrobial drug use, selection pressure, and mutation-based resistance. Thus, based on the large clonal diversity of the FQ-resistant E. coli and the resistant isolates that have a slightly elevated mutation rate relative to FQ-sensitive isolates, independent emergence of new resistant mutants likely occurs regularly in this setting.
Phylogenetic Typing and Virulence Profiling
Phylogenetic typing and virulence profiling were performed to investigate whether a potential clinical hazard was associated with the presence of these isolates. Our data on the distribution of phylogenetic groups among the 396 E. coli isolates are consistent with those of most other studies. In an examination of human commensal E. coli strains, the frequencies of B2 strains were found to be 1 (2%) of 55 in Mali, 6 (11%) of 56 in France, and 11 (19%) of 57 in Croatia (18). In our study, 4% of the isolates overall were of B2 origin. However, the results from a report by Zhang et al. do not agree with our data (19). B2 strains accounted for 42 (48%) of 88 commensal rectal strains from healthy college-aged women in Michigan. Likewise, Sannes et al. noted a high prevalence of group B2 among rectal isolates from hospitalized, elderly, male veterans in Minnesota (20). Differences may be due to geographic variation, differences in host population characteristics, or differences in strain characteristics such as antimicrobial resistance.
We did not observe a significant shift toward low-virulence phylogenetic groups for resistant isolates, as was reported by Johnson et al. (10). However, we confirmed that the isolates were notably depleted for phylogenetic group B2 and enriched for group D. We also confirmed that FQ-resistant E. coli exhibited sparser virulence profiles. The most prevalent VF was iutA, which was detected in 36% of the resistant isolates; however, this VF is less common in virulent group B2 strains (1). Accordingly, only 2% of the resistant isolates were found to be ExPEC. These 4 isolates also lacked the VFs iha, sat, fyuA, and malX as compared to the FQ-susceptible ExPEC. Whether ExPEC strains cause infection in humans depends on several other factors, including susceptibility of the host. Therefore, that many (36%) of the 44 ExPEC isolates were from group 2 patients who had been hospitalized for >5 days is of concern. When patients become colonized with FQ-resistant ExPEC strains in the hospital, they presumably will have an increased risk of acquiring a nosocomial infection and, when discharged with such a strain, also for community-acquired infection; in such case, an optimal therapy will be more difficult to select. Of note, a relationship has recently been shown to exist between ciprofloxacin-resistance in E. coli and the production of extended-spectrum β-lactamases, which would further limit therapeutic options (21).
Our observations provide insight into the epidemiology and virulence characteristics of FQ-resistant E. coli from stools of patients and healthy participants in Indonesia. The high prevalence of FQ-resistant E. coli in the hospital environment seems to be primarily due to a combination of limited clonal spread and the spontaneous emergence of resistant strains, possibly fostered by selection pressure. Transferable resistance, however, cannot be ruled out as an additional explanation in the present study and will be the subject of future investigations. Although the resistant isolates mainly belong to phylogenetic groups A and B1 and show a low virulence profile, similar strains have caused disease in humans (3,10). The data support the need to implement strict infection control measures in hospitals and to promote and monitor the prudent use of antimicrobial drugs. Continued surveillance of the changes of resistance patterns and virulence profiles of clinical and nonclinical E. coli isolates is warranted.
Dr Kuntaman is a clinical microbiologist at the Dr Soetomo Hospital in Surabaya, Indonesia, and a lecturer and researcher at the Medical Faculty of the Airlangga University, Surabaya, Indonesia. Since 1987, his research activities have focused on the mechanisms of antimicrobial resistance in Indonesia.
Acknowledgment
This work was facilitated by grant number 99-MED-3 from the Royal Netherlands Academy of Sciences and Arts in the framework of its Scientific Program Indonesia-Netherlands (SPIN), Amsterdam, the Netherlands, and by RUTI grant number RUTI II – KMRT 2003 from the Ministry of Research and Technology, Jakarta, Republic of Indonesia. The contribution of JR Johnson was supported by the Office of Research and Development, Medical Research Service, Department of Veterans Affairs; Minnesota Department of Health; and National Research Initiative Competitive Grants Program/ US Department of Agriculture grant 00-35212-9408. A van Belkum was awarded with the AstraZeneca ESCMID Turning the Tide of Resistance Grant 2003, which further supported the current study.
References
- Johnson JR, Delavari P, Kuskowski M, Stell AL. Phylogenetic distribution of extraintestinal virulence-associated traits in Escherichia coli. J Infect Dis. 2001;183:78–88. DOIPubMedGoogle Scholar
- Herzer PJ, Inouye S, Inouye M, Whittam TS. Phylogenetic distribution of branched RNA-linked multicopy single-stranded DNA among natural isolates of Escherichia coli. J Bacteriol. 1990;172:6175–81.PubMedGoogle Scholar
- Picard B, Garcia JS, Gouriou S, Duriez P, Brahimi N, Bingen E, The link between phylogeny and virulence in Escherichia coli extraintestinal infection. Infect Immun. 1999;67:546–53.PubMedGoogle Scholar
- Johnson JR, Kuskowski MA, Owens K, Gajewski A, Winokur PL. Phylogenetic origin and virulence genotype in relation to resistance to fluoroquinolones and/or extended-spectrum cephalosporins and cephamycins among Escherichia coli isolates from animals and humans. J Infect Dis. 2003;188:759–68. DOIPubMedGoogle Scholar
- Russo TA, Johnson JR. Proposal for a new inclusive designation for extraintestinal pathogenic isolates of Escherichia coli: ExPEC. J Infect Dis. 2000;181:1753–4. DOIPubMedGoogle Scholar
- Chaniotaki S, Giakouppi P, Tzouvelekis LS, Panagiotakos D, Kozanitou M, Petrikkos G, Quinolone resistance among Escherichia coli strains from community-acquired urinary tract infections in Greece. Clin Microbiol Infect. 2004;10:75–8. DOIPubMedGoogle Scholar
- Garau J, Xercavins M, Rodriguez-Carballeira M, Gomez-Vera JR, Coll I, Vidal D, Emergence and dissemination of quinolone-resistant Escherichia coli in the community. Antimicrob Agents Chemother. 1999;43:2736–41.PubMedGoogle Scholar
- Van Belkum A, Goessens W, Van der Schee C, Lemmens-den Toom N, Vos MC, Cornelissen J, Rapid emergence of ciprofloxacin-resistant enterobacteriaceae containing multiple gentamicin resistance-associated integrons in a Dutch hospital. Emerg Infect Dis. 2001;7:862–71. DOIPubMedGoogle Scholar
- Komp Lindgren P, Karlsson A, Hughes D. Mutation rate and evolution of fluoroquinolone resistance in Escherichia coli isolates from patients with urinary tract infections. Antimicrob Agents Chemother. 2003;47:3222–32. DOIPubMedGoogle Scholar
- Johnson JR, Van der Schee C, Kuskowski MA, Goessens W, Van Belkum A. Phylogenetic background and virulence profiles of fluoroquinolone-resistant clinical Escherichia coli isolates from the Netherlands. J Infect Dis. 2002;186:1852–6. DOIPubMedGoogle Scholar
- McDonald LC, Chen FJ, Lo HJ, Yin HC, Lu PL, Huang CH, Emergence of reduced susceptibility and resistance to fluoroquinolones in Escherichia coli in Taiwan and contributions of distinct selective pressures. Antimicrob Agents Chemother. 2001;45:3084–91. DOIPubMedGoogle Scholar
- Van Belkum A, Van Leeuwen W, Kluytmans J, Verbrugh H. Molecular nosocomial epidemiology: high speed typing of microbial pathogens by arbitrary primed polymerase chain reaction assays. Infect Control Hosp Epidemiol. 1995;16:658–66. DOIPubMedGoogle Scholar
- Versalovic J, Koeuth T, Lupski JR. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 1991;19:6823–31. DOIPubMedGoogle Scholar
- Clermont O, Bonacorsi S, Bingen E. Rapid and simple determination of the Escherichia coli phylogenetic group. Appl Environ Microbiol. 2000;66:4555–8. DOIPubMedGoogle Scholar
- Johnson JR, Stell AL. Extended virulence genotypes of Escherichia coli strains from patients with urosepsis in relation to phylogeny and host compromise. J Infect Dis. 2000;181:261–72. DOIPubMedGoogle Scholar
- Luria SE, Delbrück M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics. 1943;28:491–511.PubMedGoogle Scholar
- Wang M, Tran JH, Jacoby GA, Zhang Y, Wang F, Hooper DC. Plasmid-mediated quinolone resistance in clinical isolates of Escherichia coli from Shanghai, China. Antimicrob Agents Chemother. 2003;47:2242–8. DOIPubMedGoogle Scholar
- Duriez P, Clermont O, Bonacorsi S, Bingen E, Chaventre A, Elion J, Commensal Escherichia coli isolates are phylogenetically distributed among geographically distinct human populations. Microbiology. 2001;147:1671–6.PubMedGoogle Scholar
- Zhang L, Foxman B, Marrs C. Both urinary and rectal Escherichia coli isolates are dominated by strains of phylogenetic group B2. J Clin Microbiol. 2002;40:3951–5. DOIPubMedGoogle Scholar
- Sannes MR, Kuskowski MA, Owens K, Gajewski A, Johnson JR. Virulence factor profiles and phylogenetic background of Escherichia coli isolates from veterans with bacteremia versus uninfected control patients. J Infect Dis. 2004;190:2121–8. DOIPubMedGoogle Scholar
- Tolun V, Kucukbasmaci O, Torumkuney-Akbulut D, Catal C, Ang-Kucuker M, Ang O. Relationship between ciprofloxacin resistance and extended-spectrum beta-lactamase production in Escherichia coli and Klebsiella pneumoniae strains. Clin Microbiol Infect. 2004;10:72–5. DOIPubMedGoogle Scholar
Figure
Tables
Cite This Article1The first 3 authors contributed equally to this manuscript.
2Widjoseno Gardjito, Erni P. Kolopaking, Djoko Roeshadi, Eddy Rahardjo, Hari Parathon, Kuntaman Kuntaman, Ni Made Mertaniasih, Nun Zairina, Endang Isbandiati, Mariyatul Qibtiyah, Marijam Purwanta, Usman Hadi, Ariawan Soejoenoes, Budi Riyanto, Hendro Wahyono, Musrichan Adhisaputro, Bambang Triwara, Endang Sri Lestari, Bambang Wibowo, Muchlis AU Sofro, Helmia Farida, Peterhans van den Broek, Offra Duerink, Henri Verbrugh, Inge Gyssens, and Monique Keuters.
3ExPEC: papEF (P fimbrial tip pilins); papG (P adhesin); papG alleles I, II, and III; sfaS (S fimbriae); focG (F1C fimbriae); iha (putative adhesin-siderophore); bmaE (M fimbriae); gafD (G fimbriae); F17a (F17a fimbriae); clpG (CS31A adhesin); afaE8 (afimbrial adhesin VIII); fimH (type 1 fimbriae); cnf1 (cytotoxic necrotizing factor); cdtB (cytolethal distending toxin); ireA (siderophore receptor); sat (secreted autotransporter toxin); astA (S-like enterotoxin); iroN (siderophore receptor); fyuA (yersiniabactin receptor); kpsM II, K1, and K2 (kpsM II variants; group 2 capsule); kpsM III (group 3 capsule); rfc (O4 lipopolysaccharide synthesis); cvaC (colicin V); traT (serum-resistance associated outer membrane protein); ibeA (invasion of brain endothelium); ompT (outer membrane protease T); iss (increased serum survival); usp (uropathogenic specific protein); malX (marker for pathogenicity-associated island from strain CFT073); and H7 fliC variant (flagellin).
Table of Contents – Volume 11, Number 9—September 2005
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Alex van Belkum, Erasmus MC, University Medical Center Rotterdam, Department of Medical Microbiology and Infectious Diseases, Dr. Molewaterplein 40, 3015 GD, Rotterdam, the Netherlands; fax: 00-31-10-4633875
Top