Volume 13, Number 10—October 2007
Dispatch
Isolation of Bartonella sp. from Sheep Blood
Abstract
A Bartonella sp. was isolated from sheep blood. Bacterial identification was conducted by using electron microscopy and DNA sequencing of the 16S rRNA, citrate synthase, riboflavin synthase, and RNAase P genes. To our knowledge, this is the first report of ovine Bartonella infection.
Bartonella spp. are potential zoonotic pathogens that frequently cause bacteremia without overt disease in reservoir hosts (1). Several new Bartonella spp. were found in wild and domestic ruminants from Europe and North America (2–4), but previous blood cultures performed on 150 domestic sheep and 84 bighorn sheep failed to isolate Bartonella spp. (2,5).
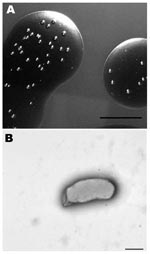
We isolated a Bartonella sp. from 2 successive lots of commercial defibrinated sheep blood received from mid- February to mid-March 2007, from a US supplier. Bottles (10 from lot 1 and 6 from lot 2), each containing 100 mL, were newly opened on the day of receipt. Approximately 0.2 mL was aseptically removed and spotted onto the surface of a Columbia agar (BBL; Becton Dickinson, Sparks, MD, USA) plate containing 5% defibrinated sheep blood. The remaining blood was stored at 4°C. Plates were incubated at 35°C in an atmosphere of 7% CO2 and examined daily for bacterial growth. Blood appeared sterile after 7 days; however, by 14–21 days, pinpoint bacterial colonies were recognized in the blood film. After 3–4 weeks, mature colonies (Figure 1, panel A) were rough, off-white, difficult to disperse but nonadherent to the agar surface, and ≈1 mm in diameter. Monomorphic colonies grew in all samples within each lot. Estimated concentrations in the starting pools of blood from lots 1 and 2 were 750 CFU/mL and 25 CFU/mL, respectively.
The cells were small, gram-negative rods, 0.47–0.60 μm in diameter, and 0.8–1.9 μm in length (Figure 1, panel B). Flagella were not observed. Growth was not detected after transfer of colonies to Columbia blood agar and several other available diagnostic media that contained blood products (e.g., chocolate agar, Centers for Disease Control and Prevention anaerobe agar, Bordet-Gengou agar, Mycoplasma agar, and hemin-supplemented thioglycolate medium). Colony transfer to Columbia blood agar plates that were first overlaid with sheep blood (from a presumed uninfected lot) resulted in only 2 or 3 colonies. Repeat cultures from lot 1 showed a 97% reduction in colony numbers after 37 days of storage and no growth in samples after 72 days of storage.
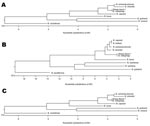
PCR was performed as described (6–8) by using template DNA obtained from a representative colony from each lot. PCR products from the lot 1 isolate (SB1) and lot 2 isolate (SB 2) had identical DNA sequences (sequencing performed at the University of Tennessee Core DNA Sequencing Facility). Phylogenetic trees of Bartonella spp. based on individual 16S rRNA, citrate synthase (gltC) and riboflavin synthase (ribC) sequence alignments showed greatest similarity to Bartonella melophagi (Figure 2).
Initial comparison of the DNA sequence with double-strand agreement from the 16S rRNA gene most closely matched (1,283/1,286 bp [99%]) that of Wolbachia melophagi (GenBank accession no. X89110). However, a shorter sequence (1,212 bp) from the SB1 isolate aligned closely (99.8%) with both W. melophagi and B. melophagi (GenBank accession nos. X89110 and AY724770). The gltC gene sequences were consistent with those of B. melophagi (GenBank accession nos. AY692475, AY724769, and AY724768). The matches of both DNA strands were 275/275 bp (100%) with each strain. The DNA sequence with double-strand agreement from the ribC gene also matched (473/473bp [100%]) that of B. melophagi (GenBank accession no. EF605287). The RNAase P gene (rnpB) sequence had more distant matching (95.6%) with sequences of B. weissi (GenBank accession no. AF376050) and B. sp. Deer 159/660/1 (95.7%) (GenBank accession no. AF376051). DNA sequences from the B. melophagi rnpB gene were not available in GenBank for comparison. The DNA sequences determined in this study have been assigned the following GenBank accession nos.: 16S rRNA (EF689897), citrate synthase (EU020109), riboflavin synthase (EU020110), and RNAase P (EU020111).
The source of Bartonella sp. was likely intrinsic contamination from bacteremia in donor sheep. Blood was obtained from multiple live sheep with sterile, closed blood collection systems and from venipuncture sites that were prepared by shearing and treatment with antiseptics. Each 5-L lot (1 L/sheep) was pooled and prepared for sale in a separate, clean, well-equipped laboratory facility. Histories of sheep were not determined. Young age (9) and contact with wildlife (2) or cross-species vectors (5) may increase the risk for Bartonella infection in sheep.
Arthropod vectors often transmit Bartonella infections. Melophagus ovinus, commonly called a sheep ked, is a hemophagous ectoparasite of sheep (5). The organism from which DNA sequence of a 16S rRNA gene was isolated was an uncultured bacterial endosymbiont of sheep keds initially called W. melophagi. However, taxonomists now agree that the organism from which the original sequence came should be removed from the genus Wolbachia and placed in the genus Bartonella (5). On the basis of DNA sequence data, candidate status was proposed for the new species B. melophagi (M. Vayssier-Taussat, L. Halos, H.-J. Bouluis, unpub. data, available from www.ncbi.nlm.gov/taxonomy/browser/wwwtaxcgi?id = 291176). An organism with DNA sequence matching that of B. melophagi was recently isolated from a sheep ked (M.Y. Kosoy, K.W. Sheff, A.I. Irkhin, unpub. data, available from www.ncbi.nlm.gov/taxonomy/browser/wwwtax.cgi?id = 291176).
Sheep blood is often used in the laboratory with the expectation that it is free of bacteria. However, routine animal health surveillance and quality control procedures may fail to detect Bartonella spp. Optimal growth conditions for this organism are unknown. In this study, growth was only observed in fresh sheep blood. We were unable to obtain sufficient growth after in vitro passage for further phenotypic characterization. A novel liquid culture medium that supported growth of Bartonella spp. also used fresh, defibrinated sheep blood as a growth supplement (10). High-level Bartonella bacteremia may be transient, and the sensitivity of PCR for detection in pooled blood or individual sheep has not been established. PCR assays performed on lots 1 and 2 after 1 month of storage did not detect Bartonella spp. PCR was not performed at the time of blood collection. Risks associated with Bartonella infection in sheep are unknown. Precautions to reduce potential transmission of Bartonella are advised when handling sheep blood.
Dr Bemis is an associate professor at the University of Tennessee College of Veterinary Medicine. His research interests include diagnostic veterinary bacteriology and mycology.
Dr Kania is an associate professor at the University of Tennessee College of Veterinary Medicine. His research interests include diagnostic veterinary immunology and molecular diagnostics.
Acknowledgment
We thank John Dunlap, Rupal Brahmbhatt, Polly Giffen, Rebekah Jones, and Mary Jean Bryant for technical assistance.
References
- Chomel BB, Boulouis HJ, Maruyama S, Breitschwerdt EB. Bartonella spp. in pets and effect on human health. Emerg Infect Dis. 2006;12:389–94.PubMedGoogle Scholar
- Chang C-C, Chomel BB, Kasten RW, Heller R, Kocan KM, Ueno H, Bartonella spp. isolated from wild and domestic ruminants in North America. Emerg Infect Dis. 2000;6:306–11. DOIPubMedGoogle Scholar
- Dehio C, Lanz C, Pohl R, Behrens P, Bermond D, Piemont Y, Bartonella schoenbuchii sp. nov., isolated from the blood of wild roe deer. Int J Syst Evol Microbiol. 2001;51:1557–65.PubMedGoogle Scholar
- Rolain JM, Rousset E, LaScola B, Duquesnel R, Raoult D. Bartonella schoenbuchensis isolated from the blood of a French cow. Ann N Y Acad Sci. 2003;990:236–8. DOIPubMedGoogle Scholar
- Halos L, Jamal T, Maillard R, Girard B, Guillot J, Chomel B, Role of Hippoboscidae flies as potential vectors of Bartonella spp. infecting wild and domestic ruminants. Appl Environ Microbiol. 2004;70:6302–5. DOIPubMedGoogle Scholar
- Johnson G, Ayers M, McClure SCC, Richardson SE, Tellier R. Detection and identification of Bartonella species pathogenic for humans by PCR amplification targeting the riboflavin synthase gene (ribC). J Clin Microbiol. 2003;41:1069–72. DOIPubMedGoogle Scholar
- Norman AF, Regnery R, Jameson P, Greene C, Krause DC. Differentiation of Bartonella-like isolates at the species level by PCR–restriction fragment length polymorphism in the citrate synthase gene. J Clin Microbiol. 1995;33:1797–803.PubMedGoogle Scholar
- Pitulle C, Strehse C, Brown JW, Breitschwerdt EB. Investigation of the phylogenetic relationships within the genus Bartonella based on comparative sequence analysis of the rnpB gene, 16S rDNA and 23S rDNA. Int J Syst Evol Microbiol. 2002;52:2075–80. DOIPubMedGoogle Scholar
- Maillard R, Grimard B, Chastant-Maillard S, Chomel B, Delcroix T, Gandoin C, Effects of cow age and pregnancy on Bartonella infection in a herd of dairy cattle. J Clin Microbiol. 2006;44:42–6. DOIPubMedGoogle Scholar
- Maggi RG, Duncan AW, Breitschwerdt EB. Novel chemically modified liquid medium that will support the growth of seven Bartonella species. J Clin Microbiol. 2005;43:2651–5. DOIPubMedGoogle Scholar
Figures
Cite This ArticleTable of Contents – Volume 13, Number 10—October 2007
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
David A. Bemis, Department of Comparative Medicine, College of Veterinary Medicine, University of Tennessee, Knoxville, TN 37996-4543, USA;
Top