Volume 17, Number 10—October 2011
Dispatch
Isolation and Phylogenetic Grouping of Equine Encephalosis Virus in Israel
Abstract
During 2008–2009 in Israel, equine encephalosis virus (EEV) caused febrile outbreaks in horses. Phylogenetic analysis of segment 10 of the virus strains showed that they form a new cluster; analysis of segment 2 showed ≈92% sequence identity to EEV-3, the reference isolate. Thus, the source of this emerging EEV remains uncertain.
Equine encephalosis is an arthropod-borne, noncontagious, febrile disease of horses. It was first described >100 years ago by A. Theiler (1) under the name equine ephemeral fever. The disease is caused by Equine encephalosis virus (EEV; genus Orbivirus: subfamily Sedoreovirinae: family Reoviridae) (2,3), which is transmitted by Culicoides spp. biting midges (4). Before 2008, EEV had been isolated only in South Africa, where 7 antigenically distinct serotypes, EEV-1–7, have been identified and characterized (3).
Orbiviruses encode at least 7 structural and 4 nonstructural (NS) proteins from 10 linear dsRNA genome segments (5). The smallest genome segment, segment 10 (Seg-10), encodes NS3, which mediates the release of virus particles from infected cells, and NS3A. The second largest of the EEV genome segments, Seg-2, encodes virus protein (VP) 2, the larger of the 2 outer-capsid proteins. By analogy with bluetongue virus (BTV), the Orbivirus type species, the virus serotype is determined by the specificity of interactions between VP2 and neutralizing antibodies generated during infection of the mammalian host. Consequently, VP2 and Seg-2 show sequence variations that correlate with serotype and, thus, can be used to determine the virus serotype (6).
From October 2008 through January 2009, a febrile horse disease that was diagnosed as equine encephalosis was observed in dozens of stables across Israel (7). The recent emergence of novel orbivirus strains (including BTV and epizootic hemorrhagic disease virus) in Europe, North America, Asia, and Australia (8) is of major concern to the worldwide livestock industry. Furthermore, the similarity of EEV to African horse sickness virus, one of the most devastating pathogens of equids, warranted further investigation of the outbreaks and molecular characterization of the virus. The molecular and sequence analyses reported here confirm the existence of EEV in Israel and identify the virus and its serotype, as well as its phylogenetic roots.
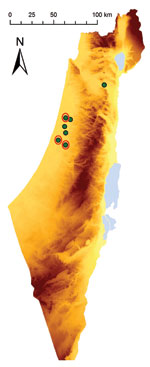
During October–November 2009, samples of whole blood from 8 febrile horses (H1–8; temperatures 39.5°C–42°C) in Israel were collected into EDTA tubes and analyzed at the Koret School of Veterinary Medicine (Hebrew University, Rehovot, Israel). Vero cell (American Type Culture Collection, Manassas, VA, USA) culture results of blood from H3, H5, and H8 were positive for EEV (Table 1; Figure 1).
Total RNA was extracted from the fifth and sixth passages of all 3 samples by using the QIAamp Viral RNA Mini Kit (QIAGEN, Valencia, CA, USA) to obtain sufficient viral load for the subsequent analyses and replications. RNA was reverse transcribed into cDNA by using the Verso cDNA Kit (Thermo Fisher Scientific, Epsom, UK). PCR amplification of the gene encoding NS3 (Seg-10) was performed on the 3 isolates by using GoTaq Green Master Mix (Promega, Madison, WI, USA) with the following primers: 5′-1GTT AAG TTT CTG CGC CAT GT23-3′, 5′-741GTA ACA CGT TTC CGC CAC G760-3′. Thermal cycling conditions for the PCR were as previously described (9); the primer annealing temperature was modified to 53.5°C. PCR products were purified by using a cDNA purification kit (ExoSAP-IT; USB, Cleveland, OH, USA), and sequencing was conducted by BigDye terminator cycle sequencing chemistry (Applied Biosystems, Foster City, CA, USA) in an ABI 3700 DNA Analyzer (Applied Biosystems) by using ABI data collection and sequence analysis software. Further analysis of the NS3 sequence was performed with Sequencer software, version 4.8 (Gene Codes Corp., Ann Arbor, MI, USA). Sequences were deposited in GenBank under accession nos. HQ441245 for H5, HQ441246 for H3, and HQ441247 for H8. The NS3 genes (Seg-10) were compared with those of different EEVs (9) and other related orbiviruses (Table 2). Phylogenic trees were generated by using the neighbor-joining and maximum likelihood methods (Phylip Inference Package version 3.68, Seqboot Program; J. Felsenstein, University of Washington, Seattle, WA, USA) to create 100 datasets (bootstrapping) and the DNA Maximum Likelihood Program version 3.5 (http://cmgm.stanford.edu/phylip/dnaml.html) to construct the trees. Finally, the Consense program version 3.5c (http://cmgm.stanford.edu/phylip/consense.html) was used to create a final consensus tree for our dataset. Broadhaven virus, a tick-borne orbivirus, was used as the outgroup in the phylogram for the gene encoding NS3.
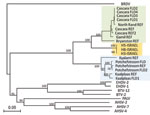
The phylogenetic analyses of EEV Seg-10 grouped the Israeli isolates with other EEV isolates but as a distinct group with no close relation to African horse sickness virus, BTV, or epizootic hemorrhagic disease virus. Within the EEV group, 3 discrete clusters (A, B, C) were recognized; the Israeli isolates formed one of these clusters (C; Figure 2). The Israeli isolates have 85%–86% nt identity to cluster A and 75%–76% nt identity to cluster B.
In addition, full-length cDNA copies of individual EEV (from H3 and H8) genome segments were synthesized and amplified by reverse transcription PCR by using the anchor spacer–ligation method as described (10,11). Partial sequences (for the upstream 450 bp) of Seg-2 from the different Israeli isolates were identical, showing 92.3% nt and 95.7% aa sequence identity with Seg-2 and VP2 of the Kaalplaas isolate, the reference isolate of EEV-3 (GenBank accession numbers are listed in Table 2). Previous phylogenetic comparisons of Seg-2/VP2 from different BTV types showed a maximum of 71% nt and 78% aa acid identity between serotypes (6), indicating that the isolates from Israel also belong to EEV type 3.
Equine encephalosis virus has long been enzootic to southern Africa, but it has not been isolated in other parts of the world. We report the characterization of an EEV strain isolated outside of Africa. Phylogenetic analysis of Seg-2 showed 92% sequence identity to EEV-3 (Kaalplaas).
Analysis of Seg-10 (the gene encoding NS3) of different orbiviruses showed 2 clusters of South African EEV strains (A and B), in agreement with previously published studies (9). These 2 clusters appear to correlate with the geographic origins of the viruses in South Africa, independent of their isolation date. It has been suggested that the 2 EEV Seg-10 clusters in South Africa are related to the distribution of their Culicoides spp. midge vectors, C. imicola (senso stricto) and C. bolitinos. The former is the most abundant Culicoides spp. midge in Israel (12). However, the EEV isolates from Israel group as a distinct cluster (C) with similar distances to the 2 South African clusters, raising questions concerning the geographic origin of this virus. A similar finding has been observed in African horse sickness virus Seg-10, which also forms into 3 distinct groups (13).
The question of how and when the virus was initially introduced to Israel remains unanswered. Because the clinical manifestations of equine encephalosis are usually mild, it is often overlooked and underdiagnosed. EEV could have been introduced to Israel before the virus was first isolated in 2009. Alternatively, the virus might have been introduced into neighboring countries and transmitted into Israel by infected vectors carried by winds, as described for other orbiviruses (14,15). The fact that the Israeli strain of EEV-3 grouped in a different cluster than the 2 South African strains, supports the idea that it has evolved in this region for a sufficient time to accumulate these changes and most likely was not recently introduced into Israel from South Africa.
Dr Aharonson-Raz is a veterinarian and a PhD candidate at the Koret School of Veterinary Medicine, Israel. Her primary research interest is epidemiology of arboviruses and infectious diseases of horses.
Acknowledgments
We thank Irit Orr for helping with the phylogenetic analysis.
Test development and analyses at Institute for Animal Health Pirbright were supported by Department for Environment, Food, and Rural Affairs, Biotechnology and Biological Sciences Research Council, and by European Union contracts OrbiVac-245266, WildTech-222633-2, and OrbiNet-K1303206.
References
- Theiler A. Notes on a fever in horses simulating horse-sickness. Transvaal Agricultural Journal. 1910;8:581–6.
- Erasmus BJ, Adelaar TF, Smit JD, Lecatsas G, Toms T. The isolation and characterization of equine encephalosis virus. Bull Off Int Epizoot. 1970;74:781–9.
- Mertens PP, Maan S, Samuel A, Attoui H. Orbivirus, Reoviridae. In: Fauquet CM, Mayo MA, Maniloff J, Desselberger U, Ball LA, editors. Virus taxonomy, VIIIth report of the ICTV. London: Elsevier/Academic Press; 2005. p. 466–83.
- Venter GJ, Groenewald DM, Paweska JT, Venter EH, Howell PG. Vector competence of selected South African Culicoides species for the Bryanston serotype of equine encephalosis virus. Med Vet Entomol. 1999;13:393–400. DOIPubMedGoogle Scholar
- Maan S, Maan NS, Samuel AR, Rao S, Attoui H, Mertens PP. Analysis and phylogenetic comparisons of full-length VP2 genes of the 24 bluetongue virus serotypes. J Gen Virol. 2007;88:621–30. DOIPubMedGoogle Scholar
- Mildenberg Z, Westcott D, Bellaiche M, Dastjerdi A, Steinbach F, Drew T. Equine encephalosis virus in Israel. Transbound Emerg Dis. 2009;56:291. DOIPubMedGoogle Scholar
- MacLachlan NJ, Guthrie AJ. Re-emergence of bluetongue, African horse sickness, and other orbivirus diseases. Vet Res. 2010;41:35. DOIPubMedGoogle Scholar
- van Niekerk M, Freeman M, Paweska JT, Howell PG, Guthrie AJ, Potgieter AC, Variation in the NS3 gene and protein in South African isolates of bluetongue and equine encephalosis viruses. J Gen Virol. 2003;84:581–90. DOIPubMedGoogle Scholar
- Maan S, Rao S, Maan NS, Anthony SJ, Attoui H, Samuel AR, Rapid cDNA synthesis and sequencing techniques for the genetic study of bluetongue and other dsRNA viruses. J Virol Methods. 2007;143:132–9. DOIPubMedGoogle Scholar
- Potgieter AC, Page NA, Liebenberg J, Wright IM, Landt O, van Dijk AA. Improved strategies for sequence-independent amplification and sequencing of viral double-stranded RNA genomes. J Gen Virol. 2009;90:1423–32. DOIPubMedGoogle Scholar
- Braverman Y, Messaddeq N, Lemble C, Kremer M. Reevaluation of the taxonomic status of the Culicoides spp. (Diptera:Ceratopogonidae) from Israel and the eastern Mediterranean and review of their potential medical and veterinary importance. J Am Mosq Control Assoc. 1996;12:437–45.PubMedGoogle Scholar
- Martin LA, Meyer AJ, O'Hara RS, Fu H, Mellor PS, Knowles NJ, Phylogenetic analysis of African horse sickness virus segment 10: sequence variation, virulence characteristics and cell exit. Arch Virol Suppl. 1998;14:281–93.PubMedGoogle Scholar
- Kedmi M, Herziger Y, Galon N, Cohen RM, Perel M, Batten C, The association of winds with the spread of EHDV in dairy cattle in Israel during an outbreak in 2006. Prev Vet Med. 2010;96:152–60. DOIPubMedGoogle Scholar
- Hendrickx G, Gilbert M, Staubach C, Elbers A, Mintiens K, Gerbier G, A wind density model to quantify the airborne spread of Culicoides species during north-western Europe bluetongue epidemic, 2006. Prev Vet Med. 2008;87:162–81. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 17, Number 10—October 2011
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Eyal Klement, Koret School of Veterinary Medicine, Robert H. Smith Faculty of Agriculture, Food and Environment, The Hebrew University, PO Box 12, Rehovot 76100, Israel
Top