Volume 17, Number 8—August 2011
Research
Enterovirus 68 among Children with Severe Acute Respiratory Infection, the Philippines
Figure 1
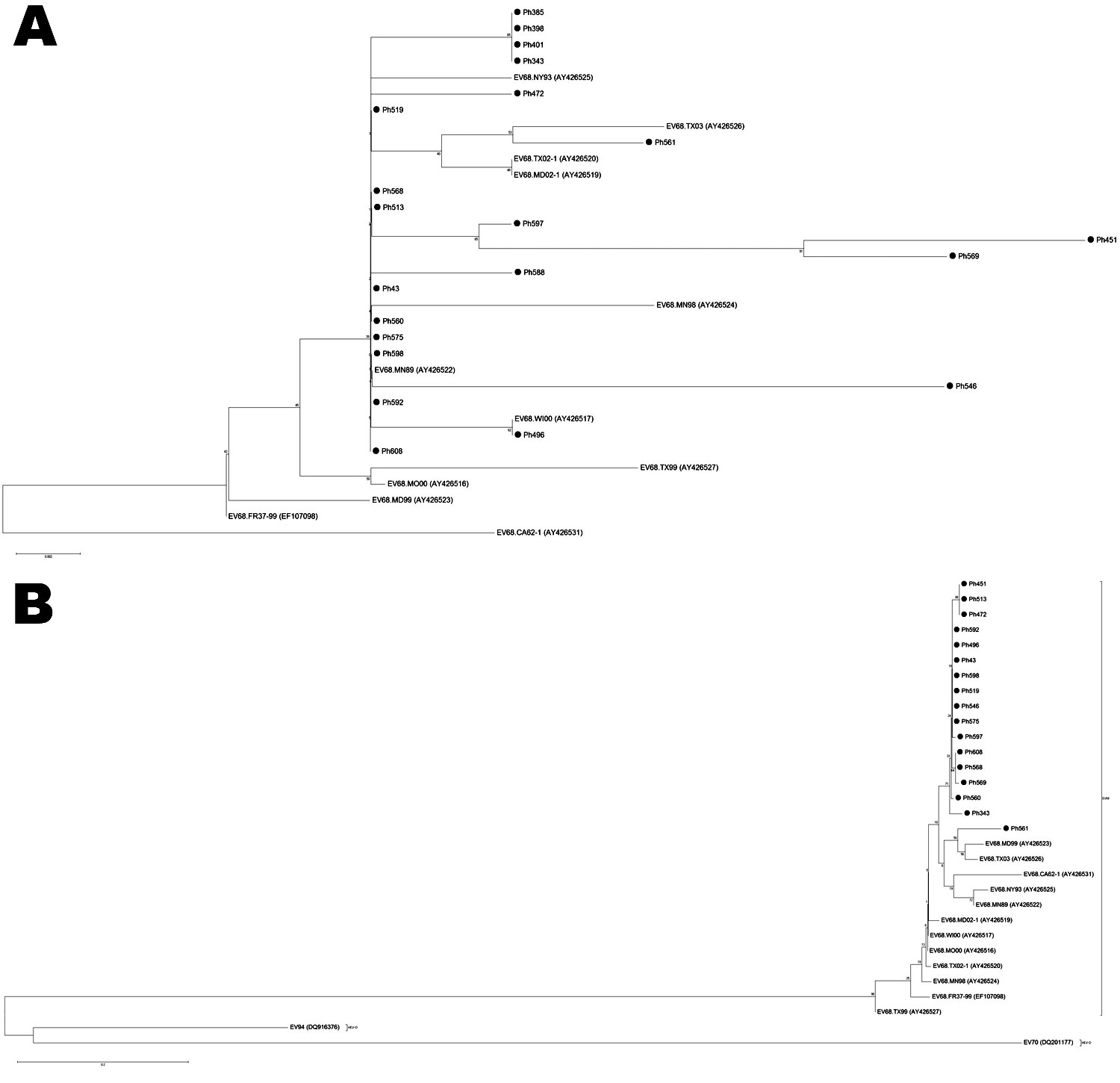
Figure 1. Phylogenetic trees of selected enterovirus (EV) 68 strains, based on the nucleotide sequence of 2 genomic regions: A) partial 5′ nontranslated region and B) partial viral protein 1. EV68 strains analyzed in this study are indicated by black circles. Phylogenetic analysis was performed by using nucleotide alignments and the neighbor-joining method, as implemented in MEGA software (www.megasoftware.net). Poliovirus 1, EV70, and EV94 sequences were used as outgroups. Scale bar indicates number of nucleotide substitutions per site.
Page created: August 15, 2011
Page updated: August 15, 2011
Page reviewed: August 15, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.