New Lineage of Lassa Virus, Togo, 2016
Shannon L.M. Whitmer
1, Thomas Strecker
1, Daniel Cadar
1, Hans-Peter Dienes, Kelly Faber, Ketan Patel, Shelley M. Brown, William G. Davis, John D. Klena, Pierre E. Rollin, Jonas Schmidt-Chanasit, Elisabeth Fichet-Calvet, Bernd Noack, Petra Emmerich, Toni Rieger, Svenja Wolff, Sarah Katharina Fehling, Markus Eickmann, Jan Philipp Mengel, Tilman Schultze, Torsten Hain, William Ampofo, Kofi Bonney, Juliana Naa Dedei Aryeequaye, Bruce Ribner, Jay B. Varkey, Aneesh K. Mehta, G. Marshall Lyon, Gerrit Kann, Philipp De Leuw, Gundolf Schuettfort, Christoph Stephan, Ulrike Wieland, Jochen W.U. Fries, Matthias Kochanek, Colleen S. Kraft, Timo Wolf, Stuart T. Nichol, Stephan Becker
2, Ute Ströher
2, and Stephan Günther
2
Author affiliations: Centers for Disease Control and Prevention, Atlanta, Georgia, USA (S.L.M. Whitmer, K. Patel, S.M. Brown, W.G. Davis, J.D. Klena, P.E. Rollin, S.T. Nichol, U. Ströher); Philipps University, Marburg, Germany (T. Strecker, S. Wolff, S.K. Fehling, M. Eickmann, S. Becker); German Center for Infection Research (DZIF), Partner sites, Hamburg, Giessen, and Marburg, Germany (T. Strecker, S. Wolff, S.K. Fehling, M. Eickmann, T. Hain, S. Becker, S. Günther); Bernhard Nocht Institute for Tropical Medicine, Hamburg (D. Cadar, J. Schmidt-Chanasit, E. Fichet-Calvet, B. Noack, P. Emmerich, T. Rieger, S. Günther); University Hospital of Cologne, Cologne, Germany (H.-P. Dienes, U. Wieland, J.W.U. Fries, M. Kochanek); Hospital of Hope, Mango, Togo (K. Faber); University of Rostock, Rostock, Germany (P. Emmerich); Justus-Liebig University Giessen, Giessen (J.P. Mengel, T. Schultze, T. Hain); Noguchi Memorial Institute for Medical Research, Accra, Ghana (W. Ampofo, K. Bonney, J.N.D. Aryeequaye); Emory University, Atlanta (B. Ribner, J.B. Varkey, A.K. Mehta, G.M. Lyon III, C.S. Kraft); University Hospital, Frankfurt/Main, Germany (G. Kann, P. De Leuw, G. Schuettfort, C. Stephan, T. Wolf)
Main Article
Figure
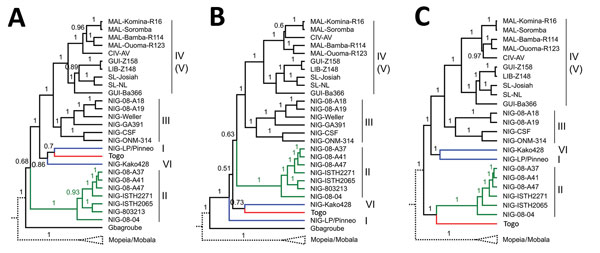
Figure. Phylogeny of the Lassa virus strain from Togo, 2016. Phylogenetic trees were inferred using BEAST2 (https://www.beast2.org/) for full-length glycoprotein precursor (A), nucleoprotein (B), and polymerase (C) genes. The analysis included representative Lassa virus strains and other Old World arenaviruses (Technical Appendix). Posterior support values are shown at the branches. Lassa virus lineages are indicated by roman numbers on the right. The branch for Mopeia and Mobala virus is shown schematically and the branches for the remaining Old World arenaviruses have been removed for clarity of presentation. The branches for the Togo strain and most closely related Lassa virus lineages are marked (red, Togo; green, lineage II; blue, lineages I and VI). The origins of the Lassa virus strains are abbreviated as follows: CIV, Côte d’Ivoire; GUI, Guinea; LIB, Liberia; MAL, Mali; NIG, Nigeria; SL, Sierra Leone.
Main Article
Page created: February 16, 2018
Page updated: February 16, 2018
Page reviewed: February 16, 2018
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
