Volume 27, Number 4—April 2021
Dispatch
Stability of SARS-CoV-2 RNA in Nonsupplemented Saliva
Abstract
The expense of saliva collection devices designed to stabilize severe acute respiratory syndrome coronavirus 2 RNA is prohibitive to mass testing. However, virus RNA in nonsupplemented saliva is stable for extended periods and at elevated temperatures. Simple plastic tubes for saliva collection will make large-scale testing and continued surveillance easier.
Despite increased diagnostic testing capacity for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), testing in many countries, including the United States, is still inadequate for slowing the coronavirus disease (COVID-19) pandemic. Many persons still do not have access to SARS-CoV-2 testing, and for some that do, an imbalance between supply and demand at large testing centers leads to long delays before results are received. The demand for testing will only increase as many schools, colleges, and workplaces reopen. Ideally, specialized population surveillance–oriented testing would require minimal diversion of resources from clinical diagnostic testing, be affordable and scalable, and enable rapid and reliable virus identification for persons with asymptomatic or subclinical infections. Thus, simplifying the sample collection and testing workflow is critical.
A simple solution is saliva collection. Saliva is a sensitive source for SARS-CoV-2 detection (1–3) and an alternative sample type for antigen and antibody testing (4,5). In addition, saliva collection is noninvasive, can be reliably performed without trained health professionals, and does not rely on a sometimes-limited swab supply. However, almost all saliva-based tests approved by the US Food and Drug Administration require specialized collection tubes containing stabilization or inactivation buffers that are costly and not always available. Moreover, as saliva continues to gain popularity as a potential specimen to aid testing demands, standardized collection methods have not been defined for saliva collection as they have for swab-based specimen collection. When true saliva is not collected (e.g., if it contains sputum), which can happen with COVID-19 inpatients when saliva is difficult to produce, specimens can be difficult to pipette (6). Combined with untested concerns regarding SARS-CoV-2 RNA stability in saliva, using supplements to reduce degradation and improve sample processing has become common. Previous work with saliva samples, however, has indicated that some buffers optimized for host nucleic acid stabilization may actually inhibit viral RNA detection (7) (S.B. Griesemer et al., unpub. data, https://doi.org/10.1101/2020.06.16.20133041), particularly in extraction-free PCRs (D.R.E. D.R.E. Ranoa et al., unpub. data, https://doi.org/10.1101/2020.06.18.159434). Thus, if true saliva (relatively easy to pipette) is being tested, the utility of collecting saliva in expensive tubes containing purported stabilization buffers comes into question. To explore the viability of broadly deploying affordable saliva-based surveillance approaches (8), we characterized SARS-CoV-2 RNA stability and virus infectivity in saliva samples stored in widely available, sterile, nuclease-free laboratory plastic (polypropylene) tubes.
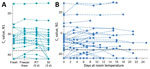
We used saliva collected from COVID-19 inpatients and at-risk healthcare workers into sterile wide-mouth containers (3) without preservatives (nonsupplemented) to evaluate the temporal stability of SARS-CoV-2 RNA at different holding temperatures (−80°C, 4°C, ≈19°C, 30°C) (Appendix). SARS-CoV-2 RNA from saliva was consistently detected at similar levels regardless of the holding time and temperatures tested. After RNA extraction and quantitative reverse transcription PCR (qRT-PCR) testing for SARS-CoV-2 on the day of saliva collection (3), we aliquoted and stored the remaining 20 sample volumes at −80°C, room temperature (≈19°C), and 30°C. Whether stored at −80°C, room temperature (5 days), or 30°C (3 days), the qRT-PCR cycle threshold (Ct) values for the N1 region of the nucleocapsid protein did not differ significantly from those for samples tested on the day of collection (Figure 1, panel A). After the freeze/thaw cycle or storage at room temperature, we observed Ct decreases of 1.058 (95% CI 2.289 to 0.141) for freeze/thaw and 0.960 (95% CI −2.219 to 0.266) for room temperature; however, the strength of this effect was low. We saw a similar effect after incubation at 30°C, with a Ct increase of 0.973 (95% CI −0.252 to 2.197). Moreover, SARS-CoV-2 RNA remained relatively stable in saliva samples left at room temperature for up to 25 days (Ct 0.027, 95% CI −0.019 to 0.071 Ct) (Figure 1, panel B). Regardless of starting Ct value (viral load), this prolonged stability of SARS-CoV-2 RNA was also observed when samples were stored for longer periods at −80°C (maximum 92 days), 4°C (maximum 21 days), and 30°C (maximum 16 days) (Appendix Figure 1).
Although SARS-CoV-2 RNA from saliva remained stable over time, we observed a decrease in human ribonuclease P at higher temperatures (room temperature, Ct 1.837, 95% CI 0.468 to 3.188 Ct; 30°C, Ct 3.526, 95% CI 1.750 to 5.349 Ct; Appendix Figure 2); the change in concentration was greater than that observed for SARS-CoV-2 RNA (Appendix Figure 3). Thus, although human RNA from saliva degrades without stabilization buffers, SARS-CoV-2 RNA remains protected even at warm temperatures suitable for nuclease activity.
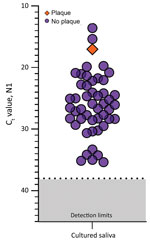
Because saliva has antiviral properties (9,10), we explored the infectiousness of SARS-CoV-2 in saliva samples. We inoculated Vero-E6 cells with 49 saliva samples with higher virus RNA titers (Ct range 13.57–35.32, median 26.01; Appendix Figure 4) because others have shown that SARS-CoV-2 isolation is uncommon from samples with low virus RNA titers (11,12; M.D. Folgueira, unpub. data, https://www.medrxiv.org/content/10.1101/2020.06.10.20127837v1). By 72 hours after inoculation, Ct values were reduced in 9 (18.7%) of the 49 cultured saliva samples tested by qRT-PCR (−12.90, −11.53, −4.30, −3.68, −3.49, −2.88, −2.81, −2.66, −2.40). Although these findings suggest an increased number of SARS-CoV-2 RNA copies by 72 hours, they may not definitively demonstrate active virus replication. For instance, Ct reductions could also result from sampling artifacts or assay variations (disparities in inoculation, RNA extraction, and qRT-PCR). To determine whether this amplification resulted from detectable, active virus replication, we performed plaque assays in triplicate with cellular lysate from 72 hours after inoculation. Only 1 of these 9 samples produced plaque-forming units; titer increased 3.79 × 104 PFU/mL at 1 hour and at 72 hours after inoculation (Figure 2). This finding suggests that increased SARS-CoV-2 genome copies identified by qRT-PCR may fall below the limit of detection in plaque assay sensitivity (100 PFU/mL) until a certain reduction in Ct is reached (e.g., Ct reduction ≤12.90) or that components of saliva possibly inhibit active viral particle production and release in vitro. A similar result has been observed when attempting to perform plaque assays of virus from the colon (13), despite studies showing that SARS-CoV-2 infects gut enterocytes (14).
The cost of commercial tubes specialized for saliva collection and SARS-CoV-2 RNA stabilization (>$7/tube) (Table) can be prohibitive for mass testing. Inexpensive saliva-based testing methods are urgently needed to help reach the capacity required to safely reopen schools and workplaces. We demonstrate the stability of SARS-CoV-2 RNA detection in saliva stored for prolonged periods in a variety of settings, which indicates that saliva can be simply collected without the need for additives.
Previous studies have demonstrated the ease with which saliva can be collected into simple, wide-mouth containers (3,15) and that buffers marketed for RNA stabilization may be detrimental to SARS-CoV-2 detection (S.B. Griesemer et al., unpub data, https://doi.org/10.1101/2020.06.16.20133041). Although some of these buffers are also marketed for virus inactivation, SARS-CoV-2 is still considered a Biosafety Level 2 hazard, meaning that with or without buffer, any saliva sample should still be handled with care. Without the need for RNA stabilization and given the limited evidence of virus replication in saliva samples, affordable alternatives to making testing accessible throughout the country are simple, sterile, nuclease-free plastic containers.
SARS-CoV-2 stability at room temperature and at 30°C permits more affordable collection and transport strategies without the need for expensive cooling strategies. Absence of the requirement for cold chain handling also makes saliva testing easier in regions with limited resources. Thus, one key for meeting mass testing demands is collection of saliva in simple, sterile, nuclease-fee tubes, negating the high costs associated with specialized collection devices.
Ms. Ott is a research assistant in the Department of Epidemiology of Microbial Diseases at the Yale School of Public Health, New Haven CT. Her work focuses on developing diagnostic tools for and analyzing functional evolution of viral pathogens, with particular focus on SARS-CoV-2 and endemic arboviruses.
Acknowledgments
We gratefully acknowledge the study participants for their time and commitment to the study. We thank all members of the clinical team at Yale New Haven Hospital for their dedication and work, which made this study possible. We also thank S. Taylor and P. Jack for technical discussions.
This study was supported by the Huffman Family Donor Advised Fund (N.D.G.), Fast Grant funding support from the Emergent Ventures at the Mercatus Center, George Mason University (N.D.G.), the Yale Institute for Global Health (N.D.G.), and the Beatrice Kleinberg Neuwirth Fund (A.I.K.). C.B.F.V. is supported by NWO Rubicon 019.181EN.004.
Yale IMPACT Research Team authors (in alphabetical order): Kelly Anastasio, Michael H. Askenase, Maria Batsu, Sean Bickerton, Kristina Brower, Molly L. Bucklin, Staci Cahill, Yiyun Cao, Edward Courchaine, Giuseppe DeIuliis, Rebecca Earnest, Renata Filler, Bertie Geng, Benjamin Goldman-Israelow, Ryan Handoko, William Khoury-Hanold, Daniel Kim, Lynda Knaggs, Maxine Kuang, Eriko Kudo, Sarah Lapidus, Joseph Lim, Melissa Linehan, Peiwen Lu, Alice Lu-Culligan, Amyn A. Malik, Anjelica Martin, Irene Matos, David McDonald, Maksym Minasyan, Nida Naushad, Jessica Nouws, Abeer Obaid, Camila Odio, Ji Eun Oh, Saad Omer, Annsea Park, Hong-Jai Park, Xiaohua Peng, Mary Petrone, Sarah Prophet, Tyler Rice, Kadi-Ann Rose, Lorenzo Sewanan, Lokesh Sharma, Denise Shepard, Mikhail Smolgovsky, Nicole Sonnert, Yvette Strong, Codruta Todeasa, Jordan Valdez, Sofia Velazquez, Arvind Venkataraman, Pavithra Vijayakumar, Elizabeth B. White, Yexin Yang.
References
- Hanson KE, Barker AP, Hillyard DR, Gilmore N, Barrett JW, Orlandi RR, et al. Self-collected anterior nasal and saliva specimens versus health care worker-collected nasopharyngeal swabs for the molecular detection of SARS-CoV-2. J Clin Microbiol. 2020;58:e01824–20. DOIPubMedGoogle Scholar
- Byrne RL, Kay GA, Kontogianni K, Aljayyoussi G, Brown L, Collins AM, et al. Saliva alternative to upper respiratory swabs for SARS-CoV-2 diagnosis. Emerg Infect Dis. 2020;26:2770–1. DOIPubMedGoogle Scholar
- Wyllie AL, Fournier J, Casanovas-Massana A, Campbell M, Tokuyama M, Vijayakumar P, et al. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N Engl J Med. 2020;383:1283–6. DOIPubMedGoogle Scholar
- Isho B, Abe KT, Zuo M, Jamal AJ, Rathod B, Wang JH, et al. Persistence of serum and saliva antibody responses to SARS-CoV-2 spike antigens in COVID-19 patients. Sci Immunol. 2020;5:
eabe5511 .PubMedGoogle Scholar - Pisanic N, Randad PR, Kruczynski K, Manabe YC, Thomas DL, Pekosz A, et al. COVID-19 serology at population scale: SARS-CoV-2-specific antibody responses in saliva. J Clin Microbiol. 2020;59:e02204–20. DOIPubMedGoogle Scholar
- Landry ML, Criscuolo J, Peaper DR. Challenges in use of saliva for detection of SARS CoV-2 RNA in symptomatic outpatients. J Clin Virol. 2020;130:
104567 . DOIPubMedGoogle Scholar - Jones TH, Muehlhauser V. Effect of handling and storage conditions and stabilizing agent on the recovery of viral RNA from oral fluid of pigs. J Virol Methods. 2014;198:26–31. DOIPubMedGoogle Scholar
- Vogels CBF, Watkins AE, Harden CA, Brackney DE, Shafer J, Wang J, et al. SalivaDirect: a simplified and flexible platform to enhance SARS-CoV-2 testing capacity. Med. 2020 Dec 26 [Epub ahead of print].
- Lieleg O, Lieleg C, Bloom J, Buck CB, Ribbeck K. Mucin biopolymers as broad-spectrum antiviral agents. Biomacromolecules. 2012;13:1724–32. DOIPubMedGoogle Scholar
- Malamud D, Abrams WR, Barber CA, Weissman D, Rehtanz M, Golub E. Antiviral activities in human saliva. Adv Dent Res. 2011;23:34–7. DOIPubMedGoogle Scholar
- Wölfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Müller MA, et al. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581:465–9. DOIPubMedGoogle Scholar
- Bullard J, Dust K, Funk D, Strong JE, Alexander D, Garnett L, et al. Predicting infectious SARS-CoV-2 from diagnostic samples. Clin Infect Dis. 2020;
ciaa638 ; [Epub ahead of print].PubMedGoogle Scholar - Zang R, Gomez Castro MF, McCune BT, Zeng Q, Rothlauf PW, Sonnek NM, et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci Immunol. 2020;5:
eabc3582 . DOIPubMedGoogle Scholar - Lamers MM, Beumer J, van der Vaart J, Knoops K, Puschhof J, Breugem TI, et al. SARS-CoV-2 productively infects human gut enterocytes. Science. 2020;369:50–4. DOIPubMedGoogle Scholar
- Wyllie AL, Chu MLJN, Schellens MHB, van Engelsdorp Gastelaars J, Jansen MD, van der Ende A, et al. Streptococcus pneumoniae in saliva of Dutch primary school children. PLoS One. 2014;9:
e102045 . DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: February 25, 2021
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.
3Team members team are listed at the end of this article.
Table of Contents – Volume 27, Number 4—April 2021
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Anne L. Wyllie and Nathan D. Grubaugh, Yale School of Public Health, 60 College St, New Haven, CT 06511, USA
Top