Volume 27, Number 5—May 2021
Research Letter
Risk for International Importations of Variant SARS-CoV-2 Originating in the United Kingdom
Abstract
A fast-spreading severe acute respiratory syndrome coronavirus 2 variant identified in the United Kingdom in December 2020 has raised international alarm. We analyzed data from 15 countries and estimated that the chance that this variant was imported into these countries by travelers from the United Kingdom by December 7 is >50%.
The United Kingdom has detected a variant of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent coronavirus disease (COVID-19), from samples initially collected in Kent on September 20 and London on September 21, 2020 (1). The variant was associated with increased transmissibility and includes deletions at amino acid sites 69 and 70 of the spike protein (2). In mid-December, the UK government tightened measures in London and southeastern England to mitigate transmission of the fast-spreading virus variant (3). On January 5, 2021, England initiated a national lockdown that included closing all schools and nonessential businesses until mid-February (4). By December 20, restrictions for travelers from the United Kingdom had been implemented by ≈40 countries (5). The new variant (501Y) has subsequently been reported worldwide, including in the United States (6), Spain, Sweden, and France, and might be spreading without detection in countries with limited virus sequencing capacity (5).
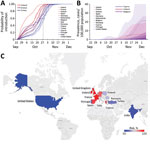
Using data from 15 countries, we estimated the probability that travelers from the United Kingdom introduced this 501Y variant into each of the countries and estimated the extent of local transmission. Our estimations were based on the changing proportion of infections caused by the 501Y variant identified in the United Kingdom (2) and population mobility from the United Kingdom to each country, determined from Facebook Data for Good (https://dataforgood.fb.com). The highest risk for importation from September 22 through December 7, 2020, was in Ireland. By October 22 (a month after the variant was first detected in the United Kingdom), the chance that 10 of the 15 countries would receive 1 imported case from the United Kingdom was at least 50% (Figure), except for Romania, Portugal, Cyprus, India, and the United States, although by November 1, this risk threshold was exceeded for all of these countries.
Using COVID-19 hospital admission data, we further estimated the local prevalence of the 501Y variant in 11 of the 15 countries, assuming that the 501Y variant is 50% more transmissible than the circulating 501N strain (Figure). The variant seems to have ascended fastest in Ireland before slowing in mid-November and is expected to be spreading rapidly in many of the other countries. As of December 7, the expected prevalence of the variant and the expected proportion of coronavirus disease cases were highest in Cyprus (prevalence 13 cases, 95% CI 0–79 cases/100,000 population; proportion 6% of cases, 95% CI 0–38% of cases) (Figure; Appendix Figures 1, 2).
These projections suggest that countries with substantial population movement from the United Kingdom were likely to harbor cases of the 501Y variant by late October 2020. Our conclusions were based on several key assumptions. The mobility data, which include ≈3 million trips from the United Kingdom to the 15 countries we analyzed, might be demographically biased by the user profile of Facebook, a major social media company with ≈2.8 billion monthly active users in the fourth quarter of 2020 (7). We assume that all introductions during this early period occurred via asymptomatic travelers from the United Kingdom and ignore possible importations from other countries or by symptomatic case-patients traveling to seek healthcare. A sensitivity analysis suggests that these assumptions may cause a downward bias in the estimated rates of global expansion (Appendix Figure 3). Furthermore, we assume a 10-day lag between infection and hospitalization on the basis of estimates from the United States (8) and Europe (9) and estimate the daily prevalence of the 501Y variant by using the method introduced in (2), under the assumptions that the 2 variants (501Y and 501N) share the same natural history (2) and symptomatic proportion (10,11). Should future studies reveal substantial epidemiologic differences between the variant and wildtype, then these estimates can be readily updated by using the full equations provided in (2).
Dr. Du is a research assistant professor in the School of Public Health, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Hong Kong Special Administrative Region, China. He develops mathematical models to elucidate the transmission dynamics, surveillance, and control of infectious diseases.
Acknowledgments
This article was preprinted at https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7814837
Financial support was provided by the Health and Medical Research Fund, Food and Health Bureau, Government of the Hong Kong Special Administrative Region (grant no. COVID190118), the US National Institutes of Health (grant no. R01 AI151176), and Centers for Disease Control and Prevention COVID Supplement (grant no. U01IP001136).
References
- Kupferschmidt K. Mutant coronavirus in the United Kingdom sets off alarms, but its importance remains unclear. 2020 Dec 20 [cited 2021 Jan 2]. https://www.sciencemag.org/news/2020/12/mutant-coronavirus-united-kingdom-sets-alarms-its-importance-remains-unclear
- Leung K, Shum MHH, Leung GM, Lam TTY, Wu JT. Early transmissibility assessment of the N501Y mutant strains of SARS-CoV-2 in the United Kingdom, October to November 2020. Euro Surveill. 2021;26:
2002106 . DOIPubMedGoogle Scholar - Kupferschmidt K. Fast-spreading U.K. virus variant raises alarms. Science. 2021;371:9–10. DOIPubMedGoogle Scholar
- Clark N, Dathan M. National lockdown – Boris Johnson orders nation to stay at home until middle of February as 13.5m Brits get Covid jab. 2021 Jan 5 [cited 2021 Jan 7]. https://www.thesun.co.uk/news/politics/13648237/national-lockdown-stay-home-boris-johnson-announcement
- BBC News. Coronavirus: Cases of new variant appear worldwide. 2020 Dec 26 [cited 2021 Jan 2]. https://www.bbc.com/news/world-europe-55452262
- Centers for Disease Control and Prevention. Emerging SARS-CoV-2 variants. 2020 Dec 29 [cited 2021 Jan 2]. https://www.cdc.gov/coronavirus/2019-ncov/more/science-and-research/scientific-brief-emerging-variants.html
- Statista. Number of monthly active Facebook users worldwide as of 4th quarter 2020 [cited 2021 Mar 20]. https://www.statista.com/statistics/264810/number-of-monthly-active-facebook-users- worldwide
- Aleta A, Martín-Corral D, Pastore Y Piontti A, Ajelli M, Litvinova M, Chinazzi M, et al. Modelling the impact of testing, contact tracing and household quarantine on second waves of COVID-19. Nat Hum Behav. 2020;4:964–71. DOIPubMedGoogle Scholar
- International Severe Acute Respiratory and Emerging Infections Consortium (ISARIC). COVID-19 report: 19 [cited 2021 Feb 1]. https://media.tghn.org/medialibrary/2020/05/ISARIC_Data_Platform_COVID-19_Report_19MAY20.pdf
- King’s College London. No evidence of change in symptoms from new coronavirus variant. King’s College London. 2021 Feb 1 [cited 2021 Mar 3]. https://www.kcl.ac.uk/news/no-evidence-change-symptoms-new-coronavirus-variant
- Davies NG, Abbott S, Barnard RC, Jarvis CI, Kucharski AJ, Munday JD, et al.; CMMID COVID-19 Working Group. COVID-19 Genomics UK (COG-UK) Consortium. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science. 2021;
eabg3055 ; [Epub ahead of print]. DOIGoogle Scholar
Figure
Cite This ArticleOriginal Publication Date: March 24, 2021
1These first authors contributed equally to this article.
Table of Contents – Volume 27, Number 5—May 2021
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Lauren Ancel Meyers, Department of Integrative Biology, The University of Texas at Austin, 2415 Speedway #C0930, Austin, TX 78712, USA
Top