Volume 9, Number 1—January 2003
Dispatch
Single Multiplex Polymerase Chain Reaction To Detect Diverse Loci Associated with Diarrheagenic Escherichia coli
Abstract
We developed and tested a single multiplex polymerase chain reaction (PCR) that detects enterotoxigenic, enteropathogenic, enteroinvasive, and Shiga-toxin–producing Escherichia coli. This PCR is specific, sensitive, and rapid in detecting target isolates in stool and food. Because of its simplicity, economy, and efficiency, this protocol warrants further evaluation in large, prospective studies of polymicrobial substances.
Escherichia coli causes disease in humans through diverse mechanisms (1). Classified on basis of their virulence traits, the most well-studied members of the diarrheagenic E. coli group include enterotoxigenic E. coli (ETEC), enteropathogenic E. coli (EPEC), enteroinvasive E. coli (EIEC), enteroaggregative E. coli (EAggEC), and Shiga-toxin–producing E. coli (STEC), also called verocytotoxin-producing or enterohemorrhagic E. coli. ETEC produce secretory toxins (enterotoxins); EPEC adhere intimately to epithelial cells and induce host cell transmembrane signaling; EIEC invade eukaryotic cells; and STEC produce Shiga toxins.
Identifying diarrheagenic E. coli in the polymicrobial milieus of stool and food poses challenges. Occasionally, economically detectable phenotypes distinguish such organisms when they are abundant in human stools. For example, sorbitol- and lactose-nonfermenting colonies are typical of E. coli O157:H7 and EIEC (2,3), respectively. However, these phenotypes are nonspecific, and subsidiary testing is needed to confirm the isolate identity. In vitro assays that detect toxins, adherence, or invasion phenotypes can also identify candidate diarrheagenic E. coli. These determinations are often expensive, require special expertise, and employ various detection systems (e.g., cell culture, cytotoxicity assays). Applying such assays to enteric microbiologic diagnosis is cumbersome.
Nucleic acid hybridization techniques, exploited by colony hybridizations or polymerase chain reaction (PCR), apply a single detection method to a diversity of organisms. The application of nucleic acid amplifications requires selecting appropriate oligonucleotide primers and optimizing conditions to maximize sensitivity and specificity. The inclusion of reactions and conditions that apply to a variety of virulence loci so that multiple candidate pathogens can be sought in a single reaction makes this technology more efficient and economical. Such multiplex detection is an appropriate solution to the challenge of finding diarrheagenic E. coli in stools and in food. We describe the development of a multiplex PCR that detects four categories of diarrheagenic E. coli and the application of the assay to human diarrheal stools and food in Mexico City.
We developed a single multiplex PCR reaction to detect ETEC, EPEC, EIEC, and STEC, using specific previously described (4–6) or new primers (GIBCO-BRL, Gaithersburg, MD) for diverse virulence traits (Table 1). Because primers for loci that unambiguously distinguish pathogenic from nonpathogenic EAggEC have not yet been determined (1), we did not address this group in this study.
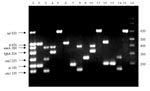
We prepared bacterial lysates by resuspending single colonies in 1 mL of deionized water (Milli-Q System, Millipore, Bedford, MA), boiling them 1 min, and then freezing them until needed. E. coli O86:H18 was the negative control in all assays. Each PCR tube contained 23 µL of reaction mix, comprised (in final concentrations) of Tris-HCl (10 mM, pH 8.3), KCl (50 mM), MgCl2 (2 mM), gelatin (100 µg/mL), glycerol (5% v/v), dATP, dCTP, dGTP, and dTTP (200 µM each), AmpliTaq polymerase (GIBCO-BRL) (0.5 U/23 µL), a mixture of the 14 primers (Table 1), and 2 µL of bacterial lysates. The final concentration of each primer in the reaction mix was determined by employing a DNA mix (Table 1) of the four prototype E. coli (7,10,11,13), until each of the seven PCR products exhibited a band of similar intensity after electrophoresis in a 2.5% agarose gel in Tris-borate-EDTA buffer and ethidium staining (Figure). The solutions were then subjected to the following cycling conditions: 50°C (2 min, 1 cycle); 95°C (5 min, 1 cycle); 95°C, 50°C, and 72°C (45 sec each temperature, 40 cycles); and a final extension step (10 min, 72°C) in a thermal cycler (iCycler System, Bio-Rad Laboratories, Inc., Hercules, CA). PCR products (4 µL) were visualized after electrophoresis and ethidium staining. The PCR sensitivity was determined by suspending one colony of each reference strain in individual 1-mL aliquots of sterile saline (0.85% w/v). Serial twofold dilutions in sterile saline were then made (to 1:256), and bacterial concentrations were determined by plating on MacConkey agar. Each dilution was also subjected to PCR analysis. E. coli 3030 (O86:H18) strain was used as a negative control during the characterization. In all further experiments, the DNA mix from the four prototype E. coli served as the positive control. The multiplex PCR was further characterized by using three additional reference strains for each category (Table 1).
Stools from 58 children <5 years of age hospitalized for diarrhea in July, August, and September, 1999, at the three main hospitals of the Instituto Mexicano del Seguro Social, Mexico City, were studied. The Institutional Review Board of the Institute approved this study, and parental informed consent was obtained for each patient. Standard diagnostic evaluations on these stools included culture for Campylobacter, Salmonella, Shigella, Vibrio cholerae, Aeromonas, and Plesiomonas; identification of Rotavirus, Adenoviridae, Astrovirus, and Caliciviridae by enzyme immunoassay; and microscopy for Entamoeba histolytica, Cryptosporidium parvum, Cyclospora cayetanensis, Isospora belli, and Giardia lamblia. Five lactose-fermenting colonies and five sorbitol-nonfermenting colonies with morphology resembling that of E. coli (when present) were selected from standard and sorbitol MacConkey agar plates, respectively, speciated biochemically, and then subjected to multiplex PCR.
Because of our concern about food safety, we purchased 52 food items (hot chili sauces and taco dressings) from street vendors in Mexico City in July, August, and September, 1999, and analyzed them for the presence of E. coli (which indicate fecal contamination) and diarrheagenic E. coli, without enrichment. One gram of food was added to 1 mL of 0.85% sterile saline and vortexed, and serial 10-fold dilutions were prepared. To enumerate candidate E. coli, and identify diarrheagenic E. coli, 100 µL of each sample and dilutions were plated on MacConkey and sorbitol MacConkey agar plates. Five pink colonies from MacConkey and five colorless colonies from sorbitol MacConkey agar were tested for indole positivity and the lactose-fermenting phenotype (if selected from the sorbitol plate). Only indole-positive, lactose-fermenting colonies isolated from both media were then subjected to the multiplex PCR. STEC from patients and food were tested to determine if they expressed the O157 lipopolysaccharide antigen by using latex particle agglutination (Oxoid Limited, Basingstoke, UK Limited, Hampshire, England).
Multiplex PCR detected the appropriate loci in each positive control strain; extraneous bands were not produced (Figure). When DNA from each of the four reference strains was mixed, the same bands appeared without nonspecific amplification (Figure). The minimum number of CFU detected were 320–1,526 for ETEC; 84–168 for EPEC; 120–1,556 for EIEC; and 20–194 for E. coli O157:H7.
Eleven (19%) of the 58 patients had candidate diarrheagenic E. coli in their stools (Table 2). In 6 (55%) of these 11 patients, no other enteric pathogens was identified, and in 3 patients target sequences were found in each of the selected E. coli colonies (Table 2). Thus, these candidate pathogens constituted the predominant aerobic coliform flora in some samples. None of the other 47 patients with diarrhea had E. coli containing the target loci in their stools. Twenty-two (42%) of the 52 food samples contained E. coli, and 7 (13%) contained candidate diarrheagenic E. coli (Table 2). No STEC isolated from patients or food expressed the O157 LPS antigen, and most were eae negative.
This multiplex PCR specifically and sensitively detected a diversity of loci in E. coli with ease, speed, and economy; its utility was demonstrated by using reference strains as well as clinical and food isolates. Conceivably, additional loci might be included because no signal attenuation occurred when a mixture of reference strains was assayed. The estimated cost per reaction for one strain is U.S. $2.00, compared to U.S. $15.00 for a colony blot analysis for one strain (data not shown). Furthermore, the signals from colony hybridizations are sometimes equivocal, in contrast to the unambiguous data obtained from our assay.
We believe that multiplex nucleic acid amplification to detect a panel of putatively pathogen traits should be considered as a replacement for tedious, less sensitive, and less specific detection technologies in clinical and food microbiologic analyses. This method should also be considered to be a more parsimonious use of PCR reagents than the individual locus PCR testing protocols described by others (15,16). Moreover, our approach does not rely on DNA extraction (16); boiling of cultures provides adequate nucleic acid to detect sequences of interest.
Comparing our protocol’s sensitivity to that reported in other protocols is difficult because of differences in methods. Specifically, other techniques seek amplicons directly from stool cultures (17) or employ fecal DNA extraction (4), whereas we assessed isolated, randomly picked colonies. Nevertheless, our sensitivity ranges were within the range of previous reports (18,19), to the extent that we were able to compare them. Our approach also provides, simultaneously, an indication of the proportion of fecal gram-negative organisms that contain loci of interest.
Without a more extensive epidemiologic analysis, we cannot state with certainty that the positive E. coli isolated were the causes of the diarrhea in the children studied. However, in some samples, the PCR-positive organisms were well represented among the aerobic coliform flora selected for analysis. Such organisms were also well represented among the food isolates. Because these E. coli indicate fecal contamination, our findings present a disconcerting picture of the hygienic status of street-vended food in Mexico City. In fact, our colony selection protocol was biased towards high-frequency organisms because we sampled only five such strains. Surveys that examine several hundred colonies (20) or PCR amplification of supernatant of fecal or food outgrowths (17,21) or of extracted DNA (4) could detect target organisms at lower densities. Though the clinical and food safety implications of low levels of candidate diarrheagenic E. coli remain unclear, multiple studies have demonstrated that consumption of food sold by street vendors is a risk factor for acquiring diarrhea in Mexico (22–24) and elsewhere (25–27), and attempts to improve the safety of these ubiquitous vehicles would most likely improve public health.
We have demonstrated for the first time that multiplex PCR can detect a variety of diarrheagenic E. coli with relative ease. Such organisms are found in food vended in Mexico City and in local children with diarrhea. This feasible technology should be evaluated in larger, controlled, prospective studies of human diarrhea and in microbiologic studies of food to establish the current epidemiology of these pathogens, including the emerging strains of STEC.
Ms. López-Saucedo is a candidate for a master of science degree in biology. Her research interests include clinical microbiology and epidemiology of diarrheal diseases.
Acknowledgment
Financial support was provided by CONAYT grants 3541-PM-9608 to FRV and I29859-M to TEG.
References
- Nataro JP, Kaper JB. Diarrheagenic Escherichia coli. Clin Microbiol Rev. 1998;11:142–201.PubMedGoogle Scholar
- Fang GD, Lima AA, Martins CV, Nataro JP, Guerrant RL. Etiology and epidemiology of persistent diarrhea in northeastern Brazil: a hospital-based, prospective, case-control study. J Pediatr Gastroenterol Nutr. 1995;21:137–44. DOIPubMedGoogle Scholar
- Flores Abuxapqui JJ, Suarez Hoil GJ, Heredia Navarrete MR, Puc Franco MA, Vivas Rosel ML. Four biochemical tests for identification of probable enteroinvasive Escherichia coli strains. Rev Latinoam Microbiol. 1999;41:259–61.PubMedGoogle Scholar
- Stacy-Phipps S, Mecca JJ, Weiss JB. Multiplex PCR assay and simple preparation method for stool specimens detect enterotoxigenic Escherichia coli DNA during the course of infection. J Clin Microbiol. 1995;33:1054–9.PubMedGoogle Scholar
- Gunzburg ST, Tornieporth NG, Riley LW. Identification of enteropathogenic Escherichia coli by PCR-based detection of the bundle-forming pilus gene. J Clin Microbiol. 1995;33:1375–7.PubMedGoogle Scholar
- Paton AW, Paton JC. Detection and characterization of Shiga toxigenic Escherichia coli by using multiplex PCR assays for stx1, stx2, eaeA, enterohemorrhagic E. coli hlyA, rfb0111, and rfb0157. J Clin Microbiol. 1998;36:598–602.PubMedGoogle Scholar
- Fleckenstein JM, Lindler LE, Elsinghorst EA, Dale JB. Identification of a gene within a pathogenicity island of enterotoxigenic Escherichia coli H10407 required for maximal secretion of the heat-labile enterotoxin. Infect Immun. 2000;68:2766–74. DOIPubMedGoogle Scholar
- Levine MM, Ristaino P, Marley G, Smith C, Knutton S, Boedeker E, Coli surface antigens 1 and 3 of colonization factor antigen II-positive enterotoxigenic Escherichia coli: morphology, purification and immune responses in humans. Infect Immun. 1984;44:409–20.PubMedGoogle Scholar
- Svennerholm AM, Wenneras C, Holmgren J, McConnell MM, Rowe B. Roles of different coli surface antigens of colonization factor antigen II in colonization by and protective immunogenicity of enterotoxigenic Escherichia coli in rabbits. Infect Immun. 1990;58:341–6.PubMedGoogle Scholar
- Okeke IN, Borneman JA, Shin S, Mellies JL, Quinn LE, Kaper JB. Comparative sequence analysis of the plasmid-encoded regulator of enteropathogenic Escherichia coli strains. Infect Immun. 2001;69:5553–64. DOIPubMedGoogle Scholar
- Schmidt H, Beutin L, Karch H. Molecular analysis of the plasmid-encoded hemolysin of Escherichia coli O157:H7 strain EDL 933. Infect Immun. 1995;63:1055–61.PubMedGoogle Scholar
- Bokete TN, Whittam TS, Wilson RA, Clausen CR, O’Callahan CM, Moseley SL, Genetic and phenotypic analysis of Escherichia coli with enteropathogenic characteristics isolated from Seattle children. J Infect Dis. 1997;175:1382–9. DOIPubMedGoogle Scholar
- Riley LW, Junio LN, Schoolnik GK. HeLa cell invasion by a strain of enteropathogenic Escherichia coli that lacks the O-antegenic polysaccharide. Mol Microbiol. 1990;4:1661–6. DOIPubMedGoogle Scholar
- Martinez MB, Whittan TS, McGraw EA, Rodrigues J, Trabulsi LR. Clonal relationship among invasive and non-invasive strains of enteroinvasive Escherichia coli serogroups. FEMS Microbiol Lett. 1999;172:145–51. DOIPubMedGoogle Scholar
- Rappelli P, Maddau G, Mannu F, Colombo MM, Fiori PL, Cappuccinelli P. Development of a set of multiplex PCR assays for the simultaneous identification of enterotoxigenic, enteropathogenic, enterohemorrhagic and enteroinvasive Escherichia coli. New Microbiol. 2001;24:77–83.PubMedGoogle Scholar
- Pass MA, Odedra R, Batt RM. Multiplex PCRs for identification of Escherichia coli virulence genes. J Clin Microbiol. 2000;38:2001–4.PubMedGoogle Scholar
- Paton AW, Paton JC, Goldwater PN, Manning PA. Direct detection of Escherichia coli Shiga-like toxin genes in primary fecal cultures by polymerase chain reaction. J Clin Microbiol. 1993;31:3063–7.PubMedGoogle Scholar
- Fratamico PM, Sackitey SK, Wiedmann M, Deng MY. Detection of Escherichia coli O157:H7 by multiplex PCR. J Clin Microbiol. 1995;33:2188–91.PubMedGoogle Scholar
- Houng HS, Sethabutr O, Echeverria P. A simple polymerase chain reaction technique to detect and differentiate Shigella and enteroinvasive Escherichia coli in human feces. Diagn Microbiol Infect Dis. 1997;28:19–25. DOIPubMedGoogle Scholar
- Samadpour M, Liston J, Ongerth JE, Tarr PI. Evaluation of DNA probes for detection of Shiga-like–toxin-producing Escherichia coli in food and calf fecal samples. Appl Environ Microbiol. 1990;56:1212–5.PubMedGoogle Scholar
- Stephan R, Ragettli S, Untermann F. Prevalence and characteristics of verotoxin-producing Escherichia coli (VTEC) in stool samples from asymptomatic human carriers working in the meat processing industry in Switzerland. J Appl Microbiol. 2000;88:335–41. DOIPubMedGoogle Scholar
- Tjoa WS, DuPont HL, Sullivan P, Pickering LK, Holguin AH, Olarte J, Location of food consumption and travelers’ diarrhea. Am J Epidemiol. 1977;106:61–6.PubMedGoogle Scholar
- Ericsson CD, Pickering LK, Sullivan P, DuPont HL. The role of location of food consumption in the prevention of travelers’ diarrhea in Mexico. Gastroenterology. 1980;79:812–6.PubMedGoogle Scholar
- Ericsson CD, DuPont HL, Mathewson JJ III. Epidemiologic observations on diarrhea developing in U.S. and Mexican students living in Guadalajara, Mexico. J Travel Med. 1995;2:6–10. DOIPubMedGoogle Scholar
- Oyemade A, Omokhodion FO, Olawuyi JF, Sridhar MK, Olaseha IO. Environmental and personal hygiene practices: risk factors for diarrhoea among children of Nigerian market women. J Diarrhoeal Dis Res. 1998;16:241–7.PubMedGoogle Scholar
- Ries AA, Vugia DJ, Beingolea L, Palacios AM, Vasquez E, Wells JG, Cholera in Piura, Peru: a modern urban epidemic. J Infect Dis. 1992;166:1429–33.PubMedGoogle Scholar
- Bryant HE, Csokonay WM, Love M, Love EJ. Self-reported illness and risk behaviours amongst Canadian travellers while abroad. Can J Public Health. 1991;82:316–9.PubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 9, Number 1—January 2003
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Teresa Estrada-García, Department of Molecular Biomedicine, CINVESTAV-IPN, Av. Instituto Politécnico Nacional 2508, Zacatenco, México D.F. 07360, México; fax: 52-555 7477134
Top