Volume 18, Number 5—May 2012
Dispatch
Human Infections with Novel Reassortant Influenza A(H3N2)v Viruses, United States, 2011
Abstract
During July–December 2011, a variant virus, influenza A(H3N2)v, caused 12 human cases of influenza. The virus contained genes originating from swine, avian, and human viruses, including the M gene from influenza A(H1N1)pdm09 virus. Influenza A(H3N2)v viruses were antigenically distinct from seasonal influenza viruses and similar to proposed vaccine virus A/Minnesota/11/2010.
Around the world, cases of human infection with swine-origin influenza viruses have been reported sporadically (1–5). From 1990 through 2010, a total of 27 cases of human infection with these viruses were confirmed by the US Centers for Disease Control and Prevention (CDC) (4,6). Of these cases, 21 were caused by triple-reassortant influenza A viruses (13 subtype H1N1, 1 subtype H1N2, and 7 subtype H3N2), which have inherited genes from classical swine, avian, and human influenza viruses. The 2009 influenza pandemic, caused by a variant triple reassortant influenza virus, influenza A(H1N1)pdm09 virus (7,8), proved that swine influenza viruses (SIVs) can cause widespread infection among humans and result in substantial economic costs. In 2010, an increase in the number of human cases of swine-origin influenza (H3N2) virus infection prompted selection of a candidate vaccine virus of swine origin, A/Minnesota/11/2010 (H3N2)v (9).
Systematic surveillance and characterization of novel viruses infecting humans and SIVs in swine are critical for early detection of viruses with pandemic potential. Since 2009, CDC has provided public health laboratories with a real-time reverse transcription PCR (rRT-PCR)–based assay for diagnostic testing for influenza (10). This assay enables detection and discrimination of influenza A virus subtypes H1N1, H3N2, and H1N1pdm09 and preliminary identification of triple-reassortant viruses possessing the nucleoprotein gene originating from SIVs.
In 2011, public health laboratories in 5 states detected 12 cases of human infection with a novel variant of influenza virus, influenza A(H3N2)v virus, by using the CDC rRT-PCR protocol. Respiratory specimens from these patients were sent to CDC for virus confirmation. History of direct or indirect contact with swine was confirmed for 6 patients. However, swine contact could not be verified for the other 6, suggesting that these infections might have been contracted through limited person-to-person transmission (11–13) (Table 1). All 12 patients recovered fully from their illness (10–12).
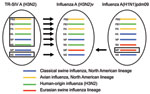
Genetic sequence analysis of RNA isolated from clinical respiratory specimens (Table 1) revealed that these influenza A(H3N2)v viruses possessed a combination of gene segments not previously found in humans (Figure 1). Of the 8 gene segments, 7 (hemagglutinin, neuraminidase, polymerase basic proteins 1 and 2, polymerase acidic protein, nucleoprotein, and nonstructural protein) were similar to those of triple-reassortant SIV A(H3N2) currently circulating in North America and to those from human triple-reassortant influenza A(H3N2) viruses isolated in 2010 from Pennsylvania, Minnesota, and Wisconsin (4), including the proposed vaccine virus of swine origin, A/Minnesota/11/2010 (14) (Figure 2, panel A; Technical Appendix Figure). However, the M genes of all 2011 influenza A(H3N2)v viruses were inherited from a pandemic (H1N1) 2009 virus (Figure 2, panel B). Although SIVs of subtypes A(H3N2) and A(H1N2) with the M gene of influenza A(H1N1)pdm09 virus have been detected in swine since 2009 (15), influenza A(H3N2)v virus possessing the M gene of influenza A(H1N1)pdm09 virus had not been detected in humans.
According to genetic analysis results, amino acid diversity among influenza A(H3N2)v hemagglutinins was low (0–3 aa) compared with that of influenza A/Minnesota/11/2010. In addition, there have been no conserved amino acid changes in the hemagglutinin when comparing 2011 influenza A(H3N2)v from humans with 2011 influenza A(H3N2) SIVs. In particular, the known receptor binding site of the hemagglutinin protein of influenza A(H3N2)v virus was typical of SIV A(H3N2) viruses recently isolated in North America.
Hemagglutinins of the influenza A(H3N2)v viruses differed substantially from the hemagglutinin of the 2011–12 human seasonal vaccine virus, A/Perth/16/2011 (58–60 aa), which resulted from divergent evolutionary paths for the H3 hemagglutinin in swine and human viruses. The effect of these substitutions on virus antigenicity was examined in the hemagglutination-inhibition assay by using a panel of reference ferret antiserum. Hemagglutination-inhibition analysis of 6 available influenza A(H3N2)v virus isolates revealed no measureable inhibition by antiserum against the current human seasonal influenza A(H3N2) vaccine virus, A/Perth/16/2009 (Table 2), indicating that influenza A(H3N2)v virus is antigenically distinct from influenza A(H3N2) viruses currently circulating among humans.
All influenza A(H3N2)v viruses tested were antigenically similar, demonstrating hemagglutination-inhibition titers with only a 2-fold difference from antiserum against other influenza A(H3N2)v viruses. These viruses were also antigenically closely related to earlier human triple-reassortant virus isolates that contained the M gene from classical SIVs (A/Wisconsin/12/2010, A/Pennsylvania/14/2010, and A/Minnesota/11/2010). All influenza A(H3N2)v viruses tested were also antigenically closely related to the proposed vaccine reassortant X-203 (13) between triple-reassortant A/Minnesota/11/2010 (H3N2) and A/PR/8/34 (H1N1) (Table 2).
The level of cross-protective immunity against influenza A(H3N2)v in humans previously vaccinated and/or exposed to previously circulated seasonal influenza A(H3N2) viruses is unknown. The antigenic characterization described here demonstrates that vaccination with the current trivalent influenza vaccine might not provide immune protection against influenza A(H3N2)v virus. A vaccine containing a contemporary influenza A(H3N2)v or an antigenically similar virus (such as A/Minnesota/11/2010) might be needed to elicit protective immunity.
Functional neuraminidase inhibition assays indicated that 6 influenza A(H3N2)v virus isolates were sensitive to the neuraminidase inhibitors oseltamivir and zanamivir. No genetic markers known to decrease sensitivity to neuraminidase inhibitors were found in the neuraminidase genes of all 12 influenza A(H3N2)v viruses. Similar to pandemic (H1N1) 2009 viruses, influenza A(H3N2)v viruses have genetic markers (V27A, S31N) in the M2 protein that confer resistance to the antiviral medications amantadine and rimantadine.
The detection of multiple cases of human infection with influenza A(H3N2)v virus within a 5-month period in 5 US states, coupled with possible human-to-human transmission, underscores the need for continued influenza surveillance at the swine–human interface. Coordinated surveillance of human and animal influenza viruses enables rapid detection of human infections with novel influenza viruses and timely identification of new virus variants in swine. As was evident during the 2009 influenza pandemic, this information is vital for development of resources that might be needed to effectively respond to the emergence and spread of a novel influenza virus in humans.
Dr Lindstrom is the team lead of the Diagnostics Development Team of the Virus Surveillance and Diagnosis Branch, Influenza Division, at the Centers for Disease Control and Prevention. His research interests are development and qualification of molecular diagnostic testing procedures for influenza viruses, confirmatory diagnostic testing and reporting of human cases of influenza, including infections potentially caused by novel influenza viruses.
Acknowledgments
We thank our collaborators from the following institutions: Pennsylvania Department of Health, Pennsylvania Department of Agriculture, Indiana State Department of Health, Indiana Board of Animal Health, Webster County Health Department, Hamilton County Public Health, Iowa Department of Public Health, University of Iowa State Hygienic Laboratory, Maine Center for Disease Control, University of Southern Maine, New Hampshire Department of Agriculture, Massachusetts Department of Agriculture, Maine Department of Agriculture, US Department of Agriculture Swine Influenza Virus Team, Mineral County Health Department, and West Virginia Department of Public Health. We also thank Thomas Gomez, Douglas Jordan, Scott Epperson, Lynette Brammer, Lyn Finelli, Susan Trock, Michael Jhung, Joseph Bresee, Michael Shaw, Daniel Jernigan, and Nancy Cox for their contributions.
Proposed vaccine reassortant X-203 was prepared in the laboratory of Doris Bucher, New York Medical School, in cooperation with the CDC Influenza Division.
References
- Myers KP, Olsen CW, Gray GC. Cases of swine influenza in humans: a review of the literature. Clin Infect Dis. 2007;44:1084–8. DOIPubMedGoogle Scholar
- Newman AP, Reisdorf E, Beinemann J, Uyeki TM, Balish A, Shu B, Human case of swine influenza A (H1N1) triple reassortant virus infection, Wisconsin. Emerg Infect Dis. 2008;14:1470–2. DOIPubMedGoogle Scholar
- Shinde V, Bridges CB, Uyeki TM, Shu B, Balish A, Xu X, Triple-reassortant swine influenza A (H1) in humans in the United States, 2005–2009. N Engl J Med. 2009;360:2616–25. DOIPubMedGoogle Scholar
- Shu B, Garten R, Emery S, Balish A, Cooper L, Sessions W, Genetic analysis and antigenic characterization of swine origin influenza viruses isolated from humans in the United States, 1990–2010. Virology. 2012;422:151–60. DOIPubMedGoogle Scholar
- Xu X, Cooper LP, Smith CB, Shu B, Deyde V, Lindstrom SL, Swine-like influenza A viruses isolated from humans from the U.S., 1990 to 2006. In: Proceedings of Options for the Control of Influenza VI; 2007 Jun 17–23; Toronto. London: International Medical Press. p. 139–141.
- Cox CM, Neises D, Garten RJ, Bryant B, Hesse RA, Anderson GA, Swine influenza virus A(H3N2) infection in human, Kansas, USA, 2009. Emerg Infect Dis. 2011;17:1143–4. DOIPubMedGoogle Scholar
- Garten RJ, Davis CT, Russell CA, Shu B, Lindstrom S, Balish A, Antigenic and genetic characteristics of swine-origin 2009 A (H1N1) influenza viruses circulating in humans. Science. 2009;325:197–201. DOIPubMedGoogle Scholar
- Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature. 2009;459:1122–5. DOIPubMedGoogle Scholar
- World Health Organization. Antigenic and genetic characteristics of influenza A(H5N1) and influenza A(H9N2) viruses and candidate vaccine viruses developed for potential use in human vaccines [cited 2012 Feb 5]. http://www.who.int/influenza/resources/documents/characteristics_virus_vaccines/en/index.html
- CDC Human Influenza Virus Real-time RT-PCR Diagnostic Panel. 2011: FDA 510K premarket notification no. K111507 [cited 2012 Feb 5]. http://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfPMN/pmn.cfm?ID=37043
- Centers for Disease Control and Prevention. Swine-origin influenza A(H3N2) virus infection in two children—Indiana and Pennsylvania, July–August 2011. MMWR Morb Mortal Wkly Rep. 2011;60:1213–5.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. Limited human-to-human transmission of novel influenza A(H3N2) virus—Iowa, November 2011. MMWR Morb Mortal Wkly Rep. 2011;60:1615–7.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. FluView Weekly Influenza Surveillance reports. 2011–2012; weeks 41, 43, 48. http://www.cdc.gov/flu/weekly/
- World Health Organization. Summary of status of development and availability of A/Minnesota/11/2010 (H3N2) swine-origin influenza virus (SOIV) candidate vaccine viruses [cited 2011 Nov 7]. http://www.who.int/influenza/vaccines/virus/candidates_reagents/a_h3n2_soiv_20111107.pdf
- Ducatez MF, Hause B, Stigger-Rosser E, Darnell D, Corzo C, Juleen K, Multiple reassortment between pandemic (H1N1) 2009 and endemic influenza viruses in pigs, United States. Emerg Infect Dis. 2011;17:1624–9. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 18, Number 5—May 2012
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Alexander Klimov, Centers for Disease Control and Prevention, Mailstop G16, 1600 Clifton Road NE, Atlanta, GA 30333, USA
Top