Volume 15, Number 6—June 2009
Research
Changes in Fluoroquinolone-Resistant Streptococcus pneumonia after 7-Valent Conjugate Vaccination, Spain
Abstract
Among 4,215 Streptococcus pneumoniae isolates obtained in Spain during 2006, 98 (2.3%) were ciprofloxacin resistant (3.6% from adults and 0.14% from children). In comparison with findings from a 2002 study, global resistance remained stable. Low-level resistance (30 isolates with MIC 4–8 μg/mL) was caused by a reserpine-sensitive efflux phenotype (n = 4) or single topoisomerase IV (parC [n = 24] or parE [n = 1]) changes. One isolate did not show reserpine-sensitive efflux or mutations. High-level resistance (68 isolates with MIC ≥16 μg/mL) was caused by changes in gyrase (gyrA) and parC or parE. New changes in parC (S80P) and gyrA (S81V, E85G) were shown to be involved in resistance by genetic transformation. Although 49 genotypes were observed, clones Spain9V-ST156 and Sweden15A-ST63 accounted for 34.7% of drug-resistant isolates. In comparison with findings from the 2002 study, clones Spain14-ST17, Spain23F-ST81, and ST8819F decreased and 4 new genotypes (ST9710A, ST57016, ST43322, and ST71733) appeared in 2006.
The bacterium Streptococcus pneumoniae is a serious cause of illness and death and a major etiologic agent of community-acquired pneumonia, meningitis, and acute otitis media. Pneumococcal resistance to antimicrobial drugs (including β-lactams, macrolides, tetracycline, and cotrimoxazole) has become a worldwide problem (1); new fluoroquinolones are being used as therapeutic alternatives for treatment of adult patients with community-acquired pneumonia (2). Resistance to fluoroquinolones in S. pneumoniae can be acquired by point mutations, intraspecific recombination (3) or interspecific recombination with the S. mitis group (3–7). Resistance is caused mainly by amino acid changes in quinolone resistance–determining regions (QRDRs) of the subunits of DNA topoisomerase IV (topo IV; parC2 and parE2) and DNA gyrase (gyrA2 and gyrB2) enzymes that control DNA topology. In addition, fluoroquinolone efflux also contributes to resistance (8). Genetic and biochemical studies have shown that for most fluoroquinolones, such as ciprofloxacin and levofloxacin, topo IV and gyrase are primary and secondary targets, respectively (9–13). However, gyrase is the primary target for moxifloxacin (14).
Although current prevalence of fluoroquinolone resistance in pneumococci is <5% (15–17), surveillance is necessary. Introduction of the 7-valent conjugate pneumococcal vaccine (PCV7), which includes serotypes such as 6B, 9V, 14, and 23F that are often associated with resistance to fluoroquinolones and other antimicrobial drugs, has resulted in changes in the epidemiology of invasive pneumococcal disease (18–20). Since the introduction of PCV7 in Spain in late 2001, ≈47% of children have been vaccinated (21).
In this study, we investigated the prevalence of fluoroquinolone-resistant pneumococci in Spain during 2006. Mutations in the QRDRs of parC, parE, and gyrA were identified, and the presence of reserpine-sensitive fluoroquinolone efflux was determined. In addition, resistance associations with other antimicrobial drugs and characteristics of drug-resistant clones were determined. To better evaluate changes in the epidemiology of resistance after the introduction of PCV7 in children, we compared our results with those of a similar study that tested isolates from 2002.
Bacterial Isolates, Serotyping, Susceptibility Testing, and Genetic Transformation
We studied 4,215 S. pneumoniae isolates from 2 hospitals (in Barcelona and San Sebastián), and a sample from 110 hospitals throughout Spain (Spanish Reference Laboratory, Madrid). Of the isolates, 2,682 were from adults, 1,400 from children, and 133 from persons whose ages were unknown. A total of 2,101 (49.9%) isolates were obtained from blood or other sterile sites; 1,055 (25%) from the lower respiratory tract; 960 (22.8%) from the upper respiratory tract, otic and conjunctival sites; and 99 (2.3%) from other sites. Isolates were confirmed as S. pneumoniae by standard methods, and serotypes were determined by the Quellung reaction. Ciprofloxacin susceptibility was determined by broth microdilution tests (Sensititer; Trek Diagnostics Inc., East Grinstead, UK) and by agar dilution according to the Clinical and Laboratory Standards Institute guidelines (22). Reserpine-sensitive fluoroquinolone efflux phenotype was determined as described (23). We performed genetic transformation as described (24) by using S. pneumoniae strains R6 and T1 (25) as receptors. For selection of transformants, we used media plates containing 1 μg/mL (R6 derivatives) or 8 μg/mL (T1 derivatives) of ciprofloxacin.
Pulsed-Field Gel Electrophoresis and Multilocus Sequence Typing
Pulsed-field gel electrophoresis (PFGE) patterns were determined by using SmaI and ApaI as described (24) and compared with 26 representative clones of the Pneumococcal Molecular Epidemiology Network (26). Isolates with patterns varying by <3 bands were considered to represent the same PFGE type (27). Multilocus sequence typing was performed as described (28) with representative isolates of PFGE types shared by >3 isolates (www.mlst.net). We analyzed selected strains representative of dominant clones from the 2002 study by multilocus sequence typing.
PCR Amplification and DNA Sequence Determination
Oligonucleotides parE398 (29) and parC152 (10) were used to amplify parE and parC QRDRs. All isolates yielded fragments of 1.6 kb, with the exception of ciprofloxacin-resistant (CipR) isolates CipR17, CipR39, CipR74, and CipR76, which yielded fragments of ≈5, 5, 5, and 7 kb, respectively. These PCR fragments were sequenced as described (24). Oligonucleotides gyrA44 and gyrA170 (29) were used to amplify and sequence gyrA QRDRs. Oligonucleotides antUP and antDOWN (4) were used to detect the ant gene.
Among the 4,215 isolates studied, 98 were CipR. Of these isolates, 30 (30.6%) showed low-level resistance (LL-CipR, MICs 4–8 μg/mL) and 68 (69.4%) high-level resistance (HL-CipR, MICs 16–128 μg/mL) (Table 1). By age group, the prevalence of CipR was 0.14% (2/1,400) among isolates from pediatric patients (<15 years of age) and 3.6% (96/2,682) among isolates from adult patients. Resistance was higher among noninvasive pneumococci (3.3%, 70/2,114) than among invasive isolates (1.3%, 28/2,101, p<0.001). The highest rate of Cip resistance was found for isolates from adults >64 years of age (Table 1). All HL-CipR isolates were from adult patients; most (53/68, 77.9%) were isolated from sputum. CipR isolates showed high rates of resistance to antimicrobial drugs. However, these rates were lower than those found in the 2002 study (Table 1).
The parC, parE, and gyrA QRDRs of the 98 CipR isolates were characterized. Most CipR isolates (93/98) showed low nucleotide sequence variations (<1%) in their QRDRs, but 5 isolates showed high variations (>4%). Four of them were in parC, parE, and gyrA, and only 1 was in gyrA. These results suggest an interspecific recombinant origin for these genes. In accordance, all isolates with recombinant parE and parC genes carried the ant gene, typical of the S. mitis group (4), as shown by PCR amplification.
Twenty-one of the 98 isolates had efflux for Cip; 3 of them also had efflux for levofloxacin (Tables 2, 3), and none had efflux for moxifloxacin. Efflux was equally distributed among LL-CipR and HL-CipR isolates. The contribution of the efflux mechanism to resistance in those isolates is unclear. Mutations not previously described that produced changes in parC (D78N, S80P, D83E), parE (I476F), and gyrA (G79A, S81V, E85G, V101I) were found in 8 isolates. To test the contribution of these changes to resistance, transformation experiments using strains R6 or T1 (as R6, parC S79F) as receptors of parC or gyrA QRDRs, respectively, were performed. The QRDRs of several independent transformants were sequenced to confirm the presence of the same mutation in the donor DNA and MICs of these transformants were determined. Although no transformation was achieved with PCR products carrying parC D78N or parE I476F, transformation to increased resistance was observed with products carrying parC S80P, gyrA S81V, and gyrA E85G changes (Table 2).
Three of these changes were accompanied by other changes known to be involved in resistance: gyrA G79A with S81F; parC D83E with S79F, and gyrA V101I with S81F. Among 5 T1 transformants obtained with a gyrA QRDR carrying G79A and S81F, 4 carried G79A and S81F and only 1 carried S81F. Because all transformants had identical Cip MICs, results suggest that G79A is not involved in drug resistance. We could not discern the role of parC D83E and gyrA S81F in resistance, given that all R6-transformants had parC D83E and S79F and all T1 transformants had gyrA V101I and S81F. However, given the contribution to resistance of the accompanied mutations, their role in resistance is unlikely.
The contribution of classical and new mutations to Cip resistance described here enabled us to classify resistant isolates (Tables 2, 3). Five LL-CipR isolates did not show changes involved in resistance in their parC, parE, or gyrA QRDRs, including 1 with recombinant genes (Table 2). Four of them showed a reserpine-sensitive efflux phenotype for Cip (Table 2) as a single mechanism of resistance. Among the remaining 25 LL-CipR isolates, 24 had mutations producing changes at parC, and 1 isolate had a single change at parE. Among 68 HL-CipR isolates, 55 (80.9%) had double changes (51 in parC and gyrA and 4 in parE and gyrA), and 13 (19.1%) had triple mutations (7 had 2 changes in parC and 1 change in gyrA; 4 had 1 change in parC, 1 change in parE, and 1 change in gyrA; 2 had 1 change in parC and 2 changes in gyrA). According to Clinical and Laboratory Standards Institute guidelines (22), only 3 of the 30 LL-CipR isolates showed intermediate resistance to levofloxacin (MIC 4 μg/mL), and the remaining 27 isolates were susceptible to levofloxacin; all were susceptible to moxifloxacin. HL-CipR isolates showed resistance (n = 66) or intermediate resistance (n = 2) to levofloxacin. Five HL-CipR isolates were susceptible to moxifloxacin, 11 showed intermediate resistance, and 52 were resistant.
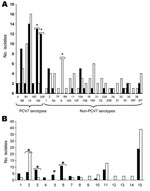
Serotype and genotype distributions of CipR isolates of 2002 (24) and 2006 were compared (Figure). Although isolates from 2006 belonged to 29 different serotypes, 5 serotypes (14, 9V, 8, 19A, and 6B) accounted for 44.9% of the total. The rate of PCV7 serotypes among CipR isolates decreased (p<0.001) in 2006 (Table 1) because of a decrease in serotypes 23F, 19F, and 6B (Figure, panel A). Forty-nine genotypes were observed among the 98 CipR isolates (Figure, panel B). Clones Spain9V-ST156 (21 isolates) and Sweden15A-ST63 (13 isolates) accounted for 34.7% of the CipR isolates. Capsular switch events were frequent in these clones (Figure): Spain9V-ST156 (12 switches) and Sweden15A-ST63 (11 switches). Four new genotypes related to non-PCV7 serotypes, (ST9710A, ST57016, ST43322, ST71733, each represented by 3 isolates) emerged in 2006 (Figure, panel B).
As we observed, isolates that shared the same PFGE pattern also shared identical polymorphisms on their DNA topoisomerase QRDRs. All but 1 of the isolates belonging to the Spain9V-ST156 clone had identical polymorphisms, the same found in the ATCC 700671 strain representative of this clone (15); the only exception was an isolate with parC, parE, and gyrA recombinant genes.
We observed a stabilization during 2002–2006 in the rates of fluoroquinolone resistance in Spain. Although the rate of Cip resistance in 2002 was 2.6% (2.2% for levofloxacin), it was 2.3% (1.7% for levofloxacin) in 2006. The rates of Cip resistance were also similar for the different age groups (3.5% for adults and 0.14% for children in 2006). However, a decrease in the rate of resistance in persons >64 years of age was found in 2006. Higher levels of resistance were found in S. pneumoniae isolated from sputa and in isolates from people >64 years of age, who more frequently have chronic obstructive pulmonary disease and who have been treated with multiple regimens of antimicrobial drugs. In accordance, development of fluoroquinolone resistance has been reported for these patients (31–33). The frequency of HL-CipR resistance in adults was 2.5% (68/2,769), slightly higher than that reported for persons in other countries in Europe (34).
Four factors may have contributed to the observed stabilization of resistance rates. These factors are fluoroquinolone use, change in circulating clones, no recommendation of fluoroquinolones for children, and fitness cost of resistance mutations.
A direct correlation between use of fluoroquinolone and prevalence of resistance in S. pneumoniae has been described (30,35). Cip use in Spain has remained stable since 1997 at 1.1 defined daily doses (DDDs)/1,000 inhabitants-days, whereas that of levofloxacin and moxifloxacin increased during 2002–2006 (from 0.2 to 0.4 DDDs/1,000 inhabitants-days for levofloxacin and from 0.3 to 0.4 DDDs/1,000 inhabitants-days for moxifloxacin, Agencia Española de Medicamentos, Madrid, Spain; http//agemed.es). Because the borderline activity of Cip against S. pneumoniae favors acquisition of first-step parC mutations (15,36), we expected that the greater activity of levofloxacin and moxifloxacin would not favor the appearance of resistance, even if one considered their increased use.
Regarding circulating pneumococcal clones, the rate of PCV7 serotypes among CipR isolates decreased from 65.3% in 2002 to 35.7% in 2006 (p<0.001). The same finding was found among CipR isolates from adults >64 years of age (7.2% in 2002 to 4.7% in 2006; p<0.02) and was probably caused by decreased transmission of pneumococci from vaccinated children to adults (37). Consequently, we have observed a decrease in 4 multidrug-resistant clones (Spain23F-ST81, Spain6B-ST90, Spain14-ST17, and ST8819F) related to PCV7-serotypes. In addition, new clones (ST6211, ST9710A, ST57016, ST43322, and ST71733) related to non-PCV7 serotypes emerged in 2006. These changes are consistent with those observed among invasive pneumococci after the introduction of PCV7 in Spain in June 2001 (38). At present, 2 clones, Spain9V-3-ST156 and Sweden15A-ST63, could be considered as the major contributors to Cip resistance in Spain, accounting for 34.7% of CipR strains.
Fluoroquinolones are not recommended for children, who are the major reservoir of pneumococci. If fluoroquinolones are given to children, according to recent reports of their safety for such use (39), increased prevalence of resistance might occur.
Regarding fitness cost of CipR mutations in S. pneumoniae, CipR isolates were divided into 3 groups. The first group is composed of 5 isolates without QRDR resistance mutations. Four isolates had a reserpine efflux phenotype. The fifth isolate may have had a different efflux inhibitor or an unknown resistance mechanism. The second group is composed of 25 LL-CipR isolates with single changes at topo IV, whose distribution, 24 at parC and 1 at parE (D435N), is consistent with the low-fitness cost of parC changes (25) and the high-fitness cost of the parE D435N change (40). The third group is composed of 68 HL-CipR isolates with gyrA changes associated with topo IV changes. GyrA changes mainly occurred at S81 (62/68), whereas changes at E85 were rare (8/68) because of the high-fitness cost of E85 changes (25).
The frequency of CipR recombinants in 2006 remained low (5.1%, 5/98 CipR isolates), similar to that in 2002 (6.7%) and that reported previously (3,4). Four isolates with mosaic parE-parC genes and long intergenic regions (4–6 kb) containing the ant gene probably originated by recombination with the S. mitis group (4). One of them belongs to the Spain9V-ST156 clone and was not typeable. The predominance of this clone and the fact that the recombinant parE-ant-parC structure did not impose a fitness cost (25) suggest recombinants could become more prevalent in the future.
Dr de la Campa is a research scientist at the Instituto de Salud Carlos III in Madrid, Spain. Her research interest focuses primarily on the molecular basis of antimicrobial drug resistance in bacteria.
Acknowledgments
Ciber Enfermedades Respiratorias is an initiative of the Instituto de Salud Carlos III.
This study was supported by grant BIO2008-02154 from Plan Nacional de I+D+I of Ministerio de Ciencia e Innovación and COMBACT-S-BIO-0260/2006 from Comunidad de Madrid.
References
- Jacobs MR, Felmingham D, Appelbaum PC, Grüneberg RN. the Alexander Project Group. The Alexander Project 1998–2000: susceptibility of pathogens isolated from community-acquired respiratory tract infection to commonly used antimicrobial agents. J Antimicrob Chemother. 2003;52:229–46. DOIPubMedGoogle Scholar
- Mandell LA, Wunderink RG, Anzueto A, Bartlett JG, Campbell GD, Dean NC, Infectious Diseases Society of America/American Thoracic Society consensus guidelines on the management of community-acquired pneumonia in adults. Clin Infect Dis. 2007;44(Suppl 2):S27–72. DOIPubMedGoogle Scholar
- Stanhope MJ, Walsh SL, Becker JA, Italia MJ, Ingraham KA, Gwynn MN, Molecular evolution perspectives on intraspecific lateral DNA transfer of topoisomerase and gyrase loci in Streptococcus pneumoniae, with implications for fluoroquinolone resistance development and spread. Antimicrob Agents Chemother. 2005;49:4315–26. DOIPubMedGoogle Scholar
- Balsalobre L, Ferrándiz MJ, Liñares J, Tubau F, de la Campa AG. Viridans group streptococci are donors in horizontal transfer of topoisomerase IV genes to Streptococcus pneumoniae. Antimicrob Agents Chemother. 2003;47:2072–81. DOIPubMedGoogle Scholar
- Bast DJ, de Azevedo JCS, Tam TY, Kilburn L, Duncan C, Mandell LA, Interspecies recombination contributes minimally to fluoroquinolone resistance in Streptococcus pneumoniae. Antimicrob Agents Chemother. 2001;45:2631–4. DOIPubMedGoogle Scholar
- Yokota S, Sato K, Kuwahara O, Habadera S, Tsukamoto N, Ohuchi H, Fluoroquinolone-resistant Streptococcus pneumoniae occurs frequently in elderly patients in Japan. Antimicrob Agents Chemother. 2002;46:3311–5. DOIPubMedGoogle Scholar
- Ferrándiz MJ, Fenoll A, Liñares J, de la Campa AG. Horizontal transfer of parC and gyrA in fluoroquinolone-resistant clinical isolates of Streptococcus pneumoniae. Antimicrob Agents Chemother. 2000;44:840–7. DOIPubMedGoogle Scholar
- Brenwald NP, Gill MJ, Wise R. Prevalence of a putative efflux mechanism among fluoroquinolone-resistant clinical isolates of Streptococcus pneumoniae. Antimicrob Agents Chemother. 1998;42:2032–5.PubMedGoogle Scholar
- Janoir C, Zeller V, Kitzis M-D, Moreau NJ, Gutmann L. High-level fluoroquinolone resistance in Streptococcus pneumoniae requires mutations in parC and gyrA. Antimicrob Agents Chemother. 1996;40:2760–4.PubMedGoogle Scholar
- Muñoz R, de la Campa AG. ParC subunit of DNA topoisomerase IV of Streptococcus pneumoniae is a primary target of fluoroquinolones and cooperates with DNA gyrase A subunit in forming resistance phenotype. Antimicrob Agents Chemother. 1996;40:2252–7.PubMedGoogle Scholar
- Tankovic J, Perichon B, Duval J, Courvalin P. Contribution of mutations in gyrA and parC genes to fluoroquinolone resistance of mutants of Streptococcus pneumoniae obtained in vivo and in vitro. Antimicrob Agents Chemother. 1996;40:2505–10.PubMedGoogle Scholar
- Pan X-S, Ambler J, Mehtar S, Fisher LM. Involvement of topoisomerase IV and DNA gyrase as ciprofloxacin targets in Streptococcus pneumoniae. Antimicrob Agents Chemother. 1996;40:2321–6.PubMedGoogle Scholar
- Fernández-Moreira E, Balas D, González I, de la Campa AG. Fluoroquinolones inhibit preferentially Streptococcus pneumoniae DNA topoisomerase IV than DNA gyrase native proteins. Microb Drug Resist. 2000;6:259–67. DOIPubMedGoogle Scholar
- Houssaye S, Gutmann L, Varon E. Topoisomerase mutations associated with in vitro selection of resistance to moxifloxacin in Streptococcus pneumoniae. Antimicrob Agents Chemother. 2002;46:2712–5. DOIPubMedGoogle Scholar
- de la Campa AG, Ferrándiz MJ, Tubau F, Pallarés R, Manresa F, Liñares J. Genetic characterization of fluoroquinolone-resistant Streptococcus pneumoniae strains isolated during ciprofloxacin therapy from a patient with bronchiectasis. Antimicrob Agents Chemother. 2003;47:1419–22. DOIPubMedGoogle Scholar
- Adam HJ, Schurek KN, Nichol KA, Hoban CJ, Baudry TJ, Laing NM, Molecular characterization of increasing fluoroquinolone resistance in Streptococcus pneumoniae isolates in Canada, 1997 to 2005. Antimicrob Agents Chemother. 2007;51:198–207. DOIPubMedGoogle Scholar
- Morrissey I, Colclough A, Northwood J. TARGETed surveillance: susceptibility of Streptococcus pneumoniae isolated from community-acquired respiratory tract infections in 2003 to fluoroquinolones and other agents. Int J Antimicrob Agents. 2007;30:345–51. DOIPubMedGoogle Scholar
- Whitney CG, Farley MM, Hadler J, Harrison LH, Bennett NM, Lynfield R, Decline in invasive pneumococcal disease after the introduction of protein-polysaccharide conjugate vaccine. N Engl J Med. 2003;348:1737–46. DOIPubMedGoogle Scholar
- Kyaw MH, Lynfield R, Schaffner W, Craig AS, Hadler J, Reingold A, Effect of introduction of the pneumococcal conjugate vaccine on drug-resistant Streptococcus pneumoniae. N Engl J Med. 2006;354:1455–63. DOIPubMedGoogle Scholar
- Beall B, McEllistrem MC, Gertz RE Jr, Wedel S, Boxrud DJ, González AL, Pre- and postvaccination clonal compositions of invasive pneumococcal serotypes for isolates collected in the United States in 1999, 2001, and 2002. J Clin Microbiol. 2006;44:999–1017. DOIPubMedGoogle Scholar
- Muñoz-Almagro C, Jordan I, Gene A, Latorre C, García-García JJ, Pallarés R. Emergence of invasive pneumococcal disease caused by nonvaccine serotypes in the era of 7-valent conjugate vaccine. Clin Infect Dis. 2008;46:174–82. DOIPubMedGoogle Scholar
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; Eighteenth informational supplement. CLSI document M100–S18. Wayne (PA): The Institute; 2008.
- Ferrándiz MJ, Oteo J, Aracil B, Gómez-Garcés JL, de la Campa AG. Drug efflux and parC mutations are involved in fluoroquinolone resistance in viridans group streptococci. Antimicrob Agents Chemother. 1999;43:2520–3.PubMedGoogle Scholar
- de la Campa AG, Balsalobre L, Ardanuy C, Fenoll A, Pérez-Trallero E, Liñares J. Fluoroquinolone resistance in penicillin-resistant Streptococcus pneumoniae clones, Spain. Emerg Infect Dis. 2004;10:1751–9.PubMedGoogle Scholar
- Balsalobre L, de la Campa AG. Fitness of Streptococcus pneumoniae fluoroquinolone-resistant strains with topoisomerase IV recombinant genes. Antimicrob Agents Chemother. 2008;52:822–30. DOIPubMedGoogle Scholar
- McGee L, McDougal L, Zhou J, Spratt BG, Tenover FC, George R, Nomenclature of major antimicrobial-resistant clones of Streptococcus pneumoniae defined by the pneumococcal molecular epidemiology network. J Clin Microbiol. 2001;39:2565–71. DOIPubMedGoogle Scholar
- Tenover FC, Arbeit R, Goering RV, Mickelsen PA, Murray BE, Persing DH, Interpreting chromosomal DNA restriction patterns produced by pulse-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–9.PubMedGoogle Scholar
- Enright MC, Spratt BG. A multilocus sequence typing scheme for Streptococcus pneumoniae: identification of clones associated with serious invasive disease. Microbiology. 1998;144:3049–60. DOIPubMedGoogle Scholar
- González I, Georgiou M, Alcaide F, Balas D, Liñares J, de la Campa AG. Fluoroquinolone resistance mutations in the parC, parE, and gyrA genes of clinical isolates of viridans group streptococci. Antimicrob Agents Chemother. 1998;42:2792–8.PubMedGoogle Scholar
- Chen DK, McGeer A, de Azavedo JC, Low DE. Decreased susceptibility of Streptococcus pneumoniae to fluoroquinolones in Canada. N Engl J Med. 1999;341:233–9. DOIPubMedGoogle Scholar
- Pérez-Trallero E, Marimón JM, Iglesias L, Larruskain J. Fluoroquinolone and macrolide treatment failure in pneumococcal pneumonia and selection of multidrug-resistant isolates. Emerg Infect Dis. 2003;9:1159–62.PubMedGoogle Scholar
- Pérez-Trallero E, Marimón JM, González A, Ercibengoa M, Larruskain J. In vivo development of high-level fluoroquinolone resistance in Streptococcus pneumoniae in chronic obstructive pulmonary disease. Clin Infect Dis. 2005;41:560–4. DOIPubMedGoogle Scholar
- de Cueto M, Rodríguez JM, Soriano MJ, López-Cerero L, Venero J, Pascual A. Fatal levofloxacin failure in treatment of a bacteremic patient infected with Streptococcus pneumoniae with a preexisting parC mutation. J Clin Microbiol. 2008;46:1558–60. DOIPubMedGoogle Scholar
- Reinert RR, Reinert S, van der Linden M, Cil MY, Al-Lahham A, Appelbaum P. Antimicrobial susceptibility of Streptococcus pneumoniae in eight European countries from 2001 to 2003. Antimicrob Agents Chemother. 2005;49:2903–13. DOIPubMedGoogle Scholar
- Liñares J, de la Campa AG, Pallarés R. Fluoroquinolone resistance in Streptococcus pneumoniae. N Engl J Med. 1999;341:1546–7. DOIPubMedGoogle Scholar
- Pérez-Trallero E, García-Arenzana JM, Jiménez JA, Peris A. Therapeutic failure and selection of resistance to quinolones in a case of pneumococcal pneumonia treated with ciprofloxacin. Eur J Clin Microbiol Infect Dis. 1990;9:905–6. DOIPubMedGoogle Scholar
- Lexau CA, Lynfield R, Danila R, Pilishvili T, Facklam R, Farley MM, Changing epidemiology of invasive pneumococcal disease among older adults in the era of pediatric pneumococcal conjugate vaccine. JAMA. 2005;294:2043–51. DOIPubMedGoogle Scholar
- Ardanuy C, Tubau F, Pallarés R, Calatayud L, Domínguez MA, Rolo D, Epidemiology of invasive pneumococcal disease among adult patients in Barcelona before and after pediatric 7-valent conjugate vaccine introduction, 1997–2007. Clin Infect Dis. 2009;48:57–64. DOIPubMedGoogle Scholar
- Murray TS, Baltimore RS. Pediatric uses of fluoroquinolone antibiotics. Pediatr Ann. 2007;36:336–42.PubMedGoogle Scholar
- Rozen DE, McGee L, Levin BR, Klugman KP. Fitness costs of fluoroquinolone resistance in Streptococcus pneumoniae. Antimicrob Agents Chemother. 2007;51:412–6. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 15, Number 6—June 2009
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Adela G. de la Campa, Unidad de Genética Bacteriana, Centro Nacional de Microbiología, Instituto de Salud Carlos III, 28220 Majadahonda, Madrid, Spain;
Top