Volume 22, Number 2—February 2016
Dispatch
Mediterranean Fin Whales (Balaenoptera physalus) Threatened by Dolphin MorbilliVirus
Abstract
During 2011–2013, dolphin morbillivirus was molecularly identified in 4 stranded fin whales from the Mediterranean Sea. Nucleoprotein, phosphoprotein, and hemagglutinin gene sequences of the identified strain were highly homologous with those of a morbillivirus that caused a 2006–2007 epidemic in the Mediterranean. Dolphin morbillivirus represents a serious threat for fin whales.
Fin whales (Balaenoptera physalus) living in the Mediterranean Sea belong to a population that is part of the Atlantic stock (1). For feeding purposes, these whales tend to concentrate in specific areas, one of which is Pelagos Sanctuary, the widest protected marine area for sea mammals in the Mediterranean Basin between Italy, France, and Monaco. In Pelagos Sanctuary and, in general, in the entire Mediterranean Sea, fin whales are considered vulnerable because of several anthropogenic threats, the most common of which are ship strikes, chemical pollution, and noise (2–4).
Postmortem investigations on well-preserved whale carcasses are necessary to gain evidence-based insight into the effect these threats have on fin whales; thus, the carcasses of all large cetaceans found stranded along the Italian coastline are systematically examined to determine the cause of death. Because morbillivirus infections have been detected during these postmortem investigations, we conducted a study to determine the effect of this natural threat on the Mediterranean fin whale population.
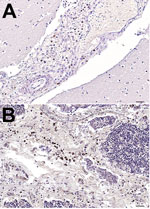
During 2006–2014, a total of 23 fin whales were found stranded along the coast of Italy. We systematically conducted full necropsies and microscopy- and molecular-based analyses on 9 (39%) carcasses that were in good conservation status. Of these 9 fin whales, 2 were juveniles and 2 were calves found during January 2011–February 2013 along the coasts of Tuscany, Sardinia, and Liguria, Italy. These young whales showed pathologic, immunohistochemical (IHC), biomolecular, and/or serologic evidence of dolphin morbillivirus (DMV) infection (Table 1) (5–9); however, not all of these cases were spatially or temporally related to known fatal DMV-associated outbreaks that occurred during 2006–2014 (Technical Appendix Figure 1). Moreover, in October 2013, a newborn male fin whale was found stranded alive on Elba Island (Tuscany); the whale died after a few hours, and postmortem investigations conducted within 24 hours of death yielded biomolecular and IHC evidence of DMV infection. Viral genome, antigen, or both were found in several tissues, along with a parasitic infection and a generalized lymphocytic depletion. Hyperimmune rabbit anti–rinderpest virus serum (provided by Pirbright Institute, Pirbright, UK) (10) was used to detect morbillivirus antigens; only circulating monocytes and tissue macrophages in brain and thymus stained positively (Figure 1).
DMV genome was detected in brain, lung, spleen, and thymus from the newborn whale; viral RNA was extracted from these tissues by using TRIzol reagent (Thermo Fisher Scientific, Waltham, MA, USA). Primer DMV2 (11) and the RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific) were used to synthesize cDNA; primer and viral RNA were incubated at 42°C for 60 min followed by 70°C for 5 min. Amplification was performed by using primers DMV-N1 and DMV-P2 (11) and Phusion Hot Start II DNA Polymerase (Thermo Scientific) with the following PCR conditions: 30 s at 98°C; 35 cycles of 10 s at 98°C, 30 s at 62°C, 1 min at 72°C; and 10 min at 72°C. Next, the DNA fragments obtained from lung and cerebral cDNA were purified, cloned into the plasmid vector pCR-Blunt II-TOPO (Invitrogen, Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instructions, and then sequenced.
Sequences from 5 lung and 4 cerebral plasmidic colonies were analyzed. Programs in the DNASTAR Lasergene software package (http://www.dnastar.com/t-dnastar-lasergene.aspx) were used to edit, assemble, and translate sequences. This technique enabled identification of a 1,355-bp DNA fragment from the newborn whale that encompassed partial nucleoprotein and phosphoprotein genes (i.e., the N1-P2 consensus fragment; GenBank accession no. KR337460). This fragment, from lung and cerebral samples, showed 98.89% sequence homology with the complete DMV genome (GenBank accession no. AJ608288) and 99.85% sequence homology with a DMV strain identified in long-finned pilot whales (Globicephala melas) that were affected by the 2006–2007 epidemic in the Mediterranean (GenBank accession no. HQ829972). Amino acid changes in KR337460, compared with AJ608288, are shown in Figure 2. The same DNA fragment was recovered from the newborn whale’s spleen and thymus by using the PCR protocol mentioned above (Technical Appendix Figure 2).
Viral hemagglutinin (H) protein mediates DMV entry into host cells by specifically binding with the whale's signaling lymphocyte activation molecule (SLAM)/CD150; thus, we also investigated variations in the H gene of the newborn whale by using the previously described cloning procedures for cerebral cDNA with 3 new overlapping primers pairs (Table 2; Figure 2). This technique enabled identification of a 1,699-bp DNA fragment encompassing a partial H gene sequence; 116 bp at the beginning of the gene were missing. The H consensus fragment, obtained from the cDNA clone for each primer pair, showed 99.41% sequence homology with the complete DMV genome (GenBank accession no. AJ608288) and 99.94% sequence homology with the DMV strain identified in long-finned pilot whales (GenBank accession no. HQ829972). We also identified 2 aa changes: the previously reported p.Val164Ala (11) and a novel variation, p.Ala451Thr, located within the H protein region (residues 382–582) involved in SLAM binding (Figure 2) (12). This variation was previously reported in other related morbilliviruses (13) and does not control any change in the tertiary structure of H antigen, as determined by using the SWISS-MODEL (http://swissmodel.expasy.org/) modeling program.
The results of our direct (IHC and biomolecular) and indirect (serologic) testing provide evidence of DMV infection or exposure in 5 (55%) of 9 fin whales that were found stranded off the Mediterranean Coast during 2011–2013. These 5 infected whales correspond to 21.7% of the 23 whales stranded along the Italian coastline during 2006–2014. The other 4 examined whales showed no evidence of morbillivirus infection. The range of DMV-susceptible host species has progressively expanded (5), as highlighted by the recent report of DMV infection in a captive common seal (Phoca vitulina) during the 2011 outbreak (14). This expansion, combined with spread of DMV through the transplacental route, resulting in virus colonization of the thymus in fetuses, could represent DMV survival strategies among cetacean populations. In addition, our data argue in favor of an epidemic cluster of fatal DMV among the Mediterranean fin whale population, even though, on the basis of the amino acid sequence of the SLAM/CD150 viral receptor, this species is not included among those susceptible to DMV epidemics (5,12).
Although the single amino acid substitution, p.Ala451Thr, did not cause substantial variations in the structure of H antigen, the effect of the variation on protein functions is unclear. Recent studies showed that similar amino acid changes could affect virulence and infectivity of different Canine distemper virus (family Paramyxoviridae, genus Morbillivirus) strains, but such changes are often neutralized by compensatory mutations that preserve the biologic activity of H protein (15). Furthermore, despite the high sequence homology observed between N, P, and H genes of the DMV strain identified in the newborn fin whale in our study and in the isolates recovered from DMV-affected cetaceans during the 1990–1992 and the 2006–2007 epidemics (GenBank accession no. AJ608288), we cannot exclude that more prominent differences occurred in virus genes encoding for both structural and nonstructural proteins responsible for virulence and pathogenicity (e.g., P/V/C and fusion genes) (5); the simultaneous occurrence of primary structure differences, if any, in the SLAM/CD150 receptor should also be taken into account (5,12). In conclusion, although further studies are needed to elucidate the complex virus–host interaction dynamics and the putative influence exerted by environmental factors, DMV should be regarded as one of the major threats for the conservation of fin whales within the Mediterranean Sea.
Dr. Mazzariol is currently assistant professor in veterinary pathology at the University of Padova, Padua, Italy. His primary research interests are diseases of marine mammals, with a special emphasis on those affecting large whales.
Acknowledgments
We thank Bruno Cozzi and Maristella Giurisato for storing and delivering samples and Giuseppe Palmisano and Emanuele Zanetti for their support during necropsies in the field.
This study was partially funded by the Italian Ministry of the Environment.
References
- Bérubé M, Aguilar A, Dendanto D, Larsen F, Notarbartolo di Sciara G, Sears R, Population genetic structure of North Atlantic, Mediterranean Sea and Sea of Cortez fin whales, Balaenoptera physalus (Linnaeus 1758): analysis of mitochondrial and nuclear loci. Mol Ecol. 1998;7:585–99. DOIPubMedGoogle Scholar
- Panigada S, Pesante G, Zanardelli M, Capoulade F, Gannier A, Weinrich MT. Mediterranean fin whales at risk from fatal ship strikes. Mar Pollut Bull. 2006;52:1287–98. DOIPubMedGoogle Scholar
- Pinzone M, Budzinski H, Tasciotti A, Ody D, Lepoint G, Schnitzler J, POPs in free-ranging pilot whales, sperm whales and fin whales from the Mediterranean Sea: influence of biological and ecological factors. Environ Res. 2015;142:185–96.PubMedGoogle Scholar
- Castellote M, Clark CW, Lammers MO. Acoustic and behavioral changes by fin whales (Balaenoptera physalus) in response to shipping and airgun noise. Biol Conserv. 2012;147:115–22. DOIGoogle Scholar
- Van Bressem MF, Duignan PJ, Banyard A, Barbieri M, Colegrove KM, De Guise S, Cetacean morbillivirus: current knowledge and future directions. Viruses. 2014;6:5145–81.PubMedGoogle Scholar
- Casalone C, Mazzariol S, Pautasso A, Di Guardo G, Di Nocera F, Lucifora G, Cetacean strandings in Italy: an unusual mortality event along the Tyrrhenian Sea coast in 2013. Dis Aquat Organ. 2014;109:81–6. DOIPubMedGoogle Scholar
- Di Guardo G, Di Francesco CE, Eleni C, Cocumelli C, Scholl F, Casalone C, Morbillivirus infection in cetaceans stranded along the Italian coastline: pathological, immunohistochemical and biomolecular findings. Res Vet Sci. 2013;94:132–7. DOIPubMedGoogle Scholar
- Mazzariol S, Marcer F, Mignone W, Serracca L, Goria M, Marsili L, Dolphin Morbillivirus and Toxoplasma gondii coinfection in a Mediterranean fin whale (Balaenoptera physalus). BMC Vet Res. 2012;8:20. DOIPubMedGoogle Scholar
- Profeta F, Di Francesco CE, Marsilio F, Mignone W, Di Nocera F, De Carlo E, Retrospective seroepidemiological investigations against Morbillivirus, Toxoplasma gondii and Brucella spp. in cetaceans stranded along the Italian coastline (1998–2014). Res Vet Sci. 2015;101:89–92. DOIPubMedGoogle Scholar
- Yang WC, Pang VF, Jeng CR, Chou LS, Chueh LL. Morbilliviral infection in a pygmy sperm whale (Kogia breviceps) from Taiwanese waters. Vet Microbiol. 2006;116:69–76. DOIPubMedGoogle Scholar
- Bellière EN, Esperón F, Sánchez-Vizcaíno JM. Genetic comparison among dolphin morbillivirus in the 1990–1992 and 2006–2008 Mediterranean outbreaks. Infect Genet Evol. 2011;11:1913–20 .PubMedGoogle Scholar
- Ohishi K, Ando A, Suzuki R, Takishita K, Kawato M, Katsumata E, Host-virus specificity of morbilliviruses predicted by structural modeling of the marine mammal SLAM, a receptor. Comp Immunol Microbiol Infect Dis. 2010;33:227–41. DOIPubMedGoogle Scholar
- Blixenkrone-Møller M, Bolt G, Jensen TD, Harder T, Svansson V. Comparative analysis of the attachment protein gene (H) of dolphin morbillivirus. Virus Res. 1996;40:47–55. DOIPubMedGoogle Scholar
- Mazzariol S, Peletto S, Mondin A, Centelleghe C, Di Guardo G, Di Francesco CE, Dolphin morbillivirus infection in a captive harbor seal (Phoca vitulina). J Clin Microbiol. 2013;51:708–11. DOIPubMedGoogle Scholar
- Sattler U, Khosravi M, Avila M, Pilo P, Langedijk JP, Ader-Ebert N, Identification of amino acid substitutions with compensational effects in the attachment protein of canine distemper virus. J Virol. 2014;88:8057–64. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 22, Number 2—February 2016
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Sandro Mazzariol, Department of Comparative Biomedicine and Food Science, University of Padova, AGRIPOLIS, Viale dell’Università 16, 35020, Legnaro (PD), Italy
Top