Volume 25, Number 1—January 2019
Research
Association of Increased Receptor-Binding Avidity of Influenza A(H9N2) Viruses with Escape from Antibody-Based Immunity and Enhanced Zoonotic Potential
Figure 1
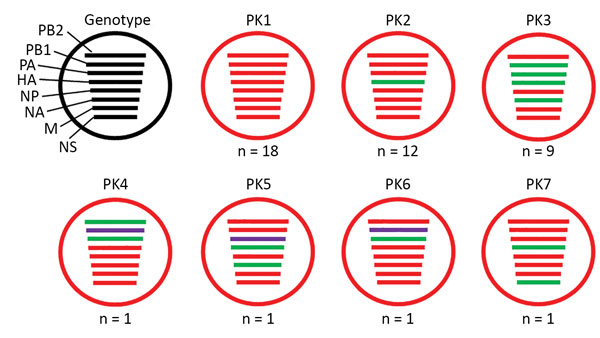
Figure 1. Genotypes of influenza A(H9N2) viruses from Pakistan. Full-genome sequences of 43 contemporary H9N2 avian influenza viruses from Pakistan were used to generate 7 unique genotypes, designated PK1–PK7. Each circle represents a genotype, and n values indicate the total number of viruses assigned to the given genotype. Each line within a circle represents a virus gene segment, and different segment colors between the same gene correspond to a >2% nucleotide difference. Black indicates wild-type virus genes; red, green, and purple indicate mutated genes. HA, hemagglutinin; M, matrix; NA, neuraminidase; NP, nucleoprotein; NS, nonstructural; PA, polymerase acidic; PB, polymerase basic.
1Current affiliation: Imperial College London, London, UK.
2Current affiliation: Chinese Academy of Agricultural Sciences, Beijing, China.